Hierarchical Part-based Generative Model for Realistic 3D Blood Vessel
Advancements in 3D vision have increased the impact of blood vessel modeling on medical applications.
Background & Academic Lineage
The problem of generating realistic 3D vascular structures emerged from the critical need for high-fidelity simulations in medical fields, such as preoperative planning and diagnostic assessment. While 3D modeling has advanced significantly, blood vessels present a unique challenge: unlike rigid objects (e.g., chairs or airplanes) that have predictable, fixed structures, vascular networks are characterized by highly irregular, branching, tree-like topologies with complex, non-uniform curvatures.
The fundamental "pain point" of previous approaches is their inability to simultaneously capture global topology and local geometric detail. Point cloud-based models struggle with the tubular, elongated nature of vessels, often failing to maintain connectivity. Meanwhile, existing generative models like VesselVAE or diffusion-based methods often treat the entire network as a single entity or lack the structural constraints necessary to prevent "block-like" artifacts or disconnected components in complex, multi-branching networks. The authors identified that previous models often failed to scale to complex datasets because they lacked a hierarchical decomposition strategy.
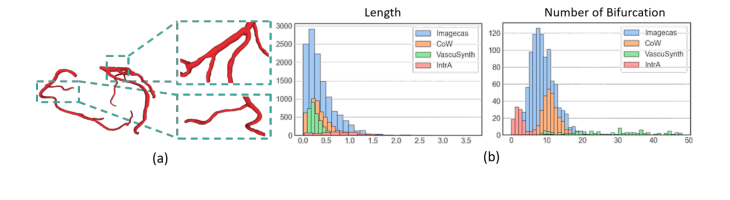 Figure 1. (a) Visualization of a Real-world Coronary Artery Dataset. The vascular net- work displays a hierarchical, tree-like structure, with localized curvatures and complex branching patterns. (b) The histograms of vessel length and number of bifurcations for four different datasets
Figure 1. (a) Visualization of a Real-world Coronary Artery Dataset. The vascular net- work displays a hierarchical, tree-like structure, with localized curvatures and complex branching patterns. (b) The histograms of vessel length and number of bifurcations for four different datasets
Intuitive Domain Terms
- Key Graph: Think of this as the "skeleton blueprint" of a tree. It ignores the thickness of the branches and focuses only on where the trunk splits and where the branches end, defining the overall layout.
- Recursive Variational Autoencoder (RVAE): Imagine a machine that learns to build a complex structure by first understanding how to assemble small, simple parts into a larger sub-assembly, and then repeating that process until the entire structure is complete.
- Geometric Descriptor: This is like a set of "instruction tags" attached to each branch, telling the model exactly how long, how curved, and how thick that specific segment should be based on its position in the overall tree.
- Implicit Neural Fields: You can think of this as a "mathematical map" that defines the shape of an object not by drawing it directly, but by creating a function that can tell you if any specific point in 3D space is "inside" or "outside" the vessel.
Notation Table
| Notation | Description |
|---|---|
| $v_{parent}$ | Attribute vector of a parent node in the key graph |
| $h_{left}, h_{right}$ | Hidden states of the left and right child nodes |
| $z_{root}$ | Global latent embedding representing the entire vascular graph |
| $C = [\ell, \delta, \kappa, \rho]$ | Geometric descriptor (length, straight-line distance, curvature, tree depth) |
| $\mathbf{x} = [x, y, z, r]$ | 3D spatial coordinates and radius of a point along a vessel segment |
| $\hat{v}, \hat{\mathbf{x}}$ | Reconstructed node attributes and segment points, respectively |
The authors solve the generation problem by decomposing it into a hierarchical, three-stage process.
-
Global Structure (Stage 1): They use an RVAE to learn the distribution of the tree topology. The encoding phase aggregates child features into a parent node via $h_{parent} = \text{MLP}(\text{concat}[v_{parent}, h_{left}, h_{right}])$. The decoding phase reverses this to generate the graph, using a classifier to predict the existence of branches. The objective is to minimize the reconstruction error of the nodes and the structural classification, regularized by a KL divergence:
$$\text{Loss} = \text{MSE}(\hat{v}, v) + \text{CrossEntropy}(\hat{y}, y) + D_{KL}(q(z_{root})\|p(z_{root}))$$ -
Local Geometry (Stage 2): Once the global structure is defined, they model individual segments as sequences. By conditioning the Transformer-based VAE on the geometric descriptor $C$, the model ensures that the generated curves match the required length and curvature defined by the key graph.
-
Assembly (Stage 3): Finally, the model performs a depth-first search traversal of the generated key graph. At each node, it applies scaling and rotation transformations to the synthesized segments to ensure they align perfectly with the global orientation $[n_x, n_y, n_z]$. This "part-based" approach effectively decouples the complex global topology from the local tubular geometry, allowing for more robust and anatomically consistent results than previous monolithic models.
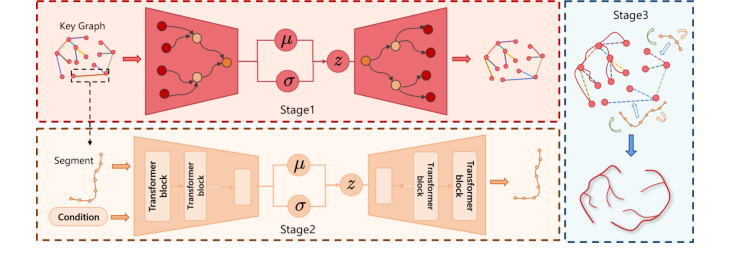 Figure 2. Overall pipeline of our method. Stage 1. Key Graph Generation: learn a global hierarchical tree. Stage 2. Vessel Segment Generation: model local 3D curve based on geometric conditions. Stage 3. Hierarchical Vessel Assembly: reconstruct the vessel skeleton by assembling segments based on the global layout
Figure 2. Overall pipeline of our method. Stage 1. Key Graph Generation: learn a global hierarchical tree. Stage 2. Vessel Segment Generation: model local 3D curve based on geometric conditions. Stage 3. Hierarchical Vessel Assembly: reconstruct the vessel skeleton by assembling segments based on the global layout
Problem Definition & Constraints
Core Problem Formulation & The Dilemma
The Starting Point (Input): The researchers begin with raw 3D medical imaging data (e.g., CCTA scans). Through preprocessing, they extract the skeleton of the vascular network—a simplified, one-dimensional representation of the vessel centerlines—along with radius information.
The Desired Endpoint (Output): The goal is to generate a high-fidelity, realistic 3D vascular model that preserves both the global topological structure (the branching tree) and the local geometric details (the specific curvature, radius, and length of individual vessel segments).
The Missing Link: Previous methods often treat the vascular network as a monolithic entity. Point cloud-based models fail to capture the tubular, elongated nature of vessels, often resulting in "holes" or disconnected components. Conversely, existing graph-based generative models often struggle to balance the global tree structure with the fine-grained, local geometric variations of individual branches. The gap lies in the inability to decouple the "where" (global topology) from the "how" (local geometry) effectively.
The Dilemma: The fundamental trade-off is between structural coherence and geometric fidelity. If a model focuses too heavily on the global tree structure, it often ignores the subtle, non-uniform curvatures and varying radii that make a vessel look "real." If it focuses too much on local point-level details, it loses the global connectivity, leading to anatomically impossible, fragmented structures.
The Harsh Constraints:
1. Topological Complexity: Blood vessels are not rigid objects; they are highly irregular, branching structures where the number and location of bifurcations vary significantly between individuals.
2. Data Sparsity & Discreteness: Standard 3D generative models (like those for chairs or airplanes) are ill-suited for the tubular, thin, and elongated nature of vessels.
3. Implicit Representation Limits: Using implicit neural fields (like in some diffusion models) often results in poor structural accuracy, as these models struggle to explicitly enforce the strict, tree-like constraints required for biological vasculature.
Why This Approach
The authors of this paper identified that traditional generative models—such as standard point cloud generators, basic Diffusion models, and VAEs—are fundamentally ill-equipped to handle the unique topological and geometric constraints of 3D vascular networks. The "inevitability" of their hierarchical part-based approach stems from the realization that blood vessels are not merely unstructured point clouds or simple volumes, but are instead complex, tree-like graphs where global connectivity and local tubular geometry are equally critical.
The Failure of Traditional SOTA
The authors explicitly reject standard "SOTA" approaches based on the following observations:
* Point Cloud-based Models: These methods treat 3D objects as unordered sets of points. While effective for rigid objects like chairs or airplanes, they fail to capture the elongated, tubular, and highly connected nature of vessels. They often produce "holes" or disconnected components because they lack an explicit understanding of the underlying skeleton.
* Implicit Neural Fields (INRs) and Diffusion: While powerful, these models often struggle with the high-dimensional noise inherent in complex branching structures. The authors note that these methods often produce "block-like" shapes or structural anomalies, failing to maintain the precise, thin-walled continuity required for medical-grade vascular simulation.
* VesselVAE: While this method attempts to use skeletal graphs, it generates the entire network as a monolithic entity. This approach lacks the modularity to handle the vast diversity of branching patterns found in real-world datasets like ImageCAS, leading to a decline in fidelity as the number of bifurcations increases.
Comparative Superiority: The Structural Advantage
The proposed method is qualitatively superior because it enforces a hierarchical decomposition that aligns with the biological reality of vasculature:
1. Global-Local Decoupling: By separating the global binary tree (the "key graph") from the local geometric details (the "segments"), the model reduces the complexity of the generation task. Instead of trying to learn the entire 3D structure at once, the model learns a high-level topological map first, then fills in the details.
2. Constraint Alignment: The "marriage" between the problem and the solution is found in the use of a Recursive Variational Autoencoder (RVAE) for the global structure and a Transformer-based VAE for the local segments. The RVAE perfectly captures the tree-like hierarchy, while the Transformer is uniquely suited to model the sequential nature of tubular curves.
3. Geometric Conditioning: The introduction of the geometric descriptor $C = [\ell, \delta, \kappa, \rho]$ acts as a bridge between the global and local stages. By conditioning the local segment generation on these specific parameters (length, straight-line distance, curvature, and tree depth), the model ensures that each segment is not just a random curve, but one that is anatomically consistent with its position in the broader vascular tree.
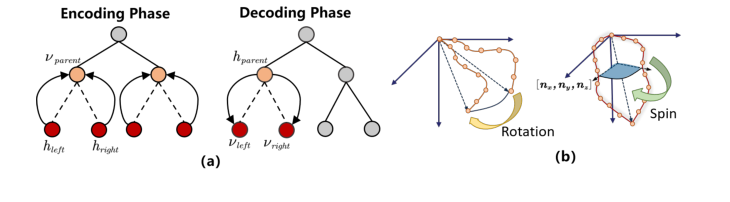 Figure 3. (a) The encoding and decoding process of the model in Stage 1. (b) The two types of rotation processes in Stage 3
Figure 3. (a) The encoding and decoding process of the model in Stage 1. (b) The two types of rotation processes in Stage 3
Mathematical & Logical Mechanism
This paper introduces a hierarchical, part-based generative framework designed to model the complex, tree-like topology and local geometry of 3D blood vessels. Unlike standard 3D generative models that treat objects as monolithic point clouds or implicit fields, this approach decomposes the vessel into a global "key graph" (the branching skeleton) and local "segments" (the tubular curves), which are then synthesized and assembled.
The Mathematical Engine
The core of the framework relies on a Recursive Variational Autoencoder (RVAE) to generate the global structure. The objective function for this stage is:
$$\text{Loss} = \text{MSE}(\hat{v}, v) + \text{CrossEntropy}(\hat{y}, y) + D_{KL}(q(z_{root}) \| p(z_{root}))$$
Tearing the Equation Apart
- $\text{MSE}(\hat{v}, v)$: This is the Mean Squared Error between the predicted node attributes $\hat{v}$ and the ground truth $v$. It acts as a geometric anchor, ensuring that the spatial coordinates and directional vectors of the generated skeleton match the real-world data.
- $\text{CrossEntropy}(\hat{y}, y)$: This term measures the classification error for the existence of child nodes. It is a logical constraint that forces the model to learn the correct branching topology (i.e., whether a vessel segment should bifurcate or terminate).
- $D_{KL}(q(z_{root}) \| p(z_{root}))$: This is the Kullback-Leibler divergence. It acts as a regularizer, forcing the latent space of the root node $z_{root}$ to follow a prior distribution (usually a Gaussian). This ensures the latent space is smooth and continuous, allowing for meaningful interpolation between different vascular structures.
Step-by-Step Flow
- Encoding: The process begins at the leaf nodes of the vessel skeleton. The model aggregates child node features into their parent using an MLP, as shown in $h_{parent} = \text{MLP}(\text{concat}[v_{parent}, h_{left}, h_{right}])$. This propagates local geometric information upward until the entire tree is compressed into a single global latent vector, $z_{root}$.
- Decoding: The process reverses. Starting from $z_{root}$, the model uses a classifier to decide if a node has children. If it does, it predicts the attributes of the child node ($\hat{v}_{left}$) and updates the hidden state to continue the recursion.
- Assembly: Once the key graph is generated, the model enters Stage 2, where a Transformer-based VAE generates the specific 3D curve for each segment, conditioned on the geometric descriptor $C$. Finally, these segments are scaled, rotated, and translated to align with the key graph, forming a complete, continuous 3D skeleton.
Results, Limitations & Conclusion
Experimental Validation
The authors "ruthlessly" tested their model against three baseline "victims": a state-of-the-art point cloud generator, TreeDiffusion, and VesselVAE.
* The Evidence: The authors used both point-based metrics (JSD, CD) and graph-based metrics (Degree distribution, Laplacian spectrum, and Graph Wasserstein Distance).
* The Result: While point-based models like PointDiffusion showed strong reconstruction metrics, they failed to maintain the topological integrity of the vessels, often producing disconnected, blocky, or "holey" meshes. The proposed model consistently achieved superior performance in graph-based metrics, proving that their part-based approach is significantly better at preserving the anatomical continuity of vascular networks.
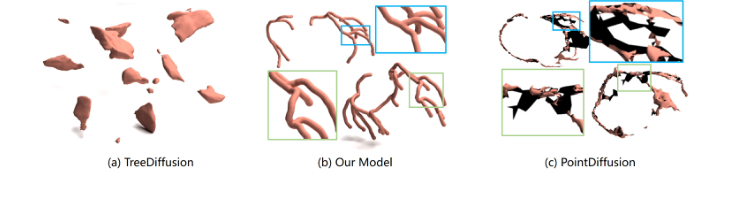 Figure 4. Reconstruction result from three different methods on ImageCAS dataset. Our approach produces more robust and anatomically consistent results compared to point cloud-based and INR-based methods
Figure 4. Reconstruction result from three different methods on ImageCAS dataset. Our approach produces more robust and anatomically consistent results compared to point cloud-based and INR-based methods
Future Discussion Topics
- Dynamic Vasculature: The current model focuses on static structures. How could this framework be extended to model the pulsatile nature of blood vessels or the dynamic changes in vascular networks during disease progression?
- Integration with Fluid Dynamics: Since this model generates highly realistic, anatomically consistent skeletons, could it be used as a prior to accelerate Computational Fluid Dynamics (CFD) simulations?
- Cross-Domain Applicability: The hierarchical part-based approach seems highly transferable. Could this architecture be adapted to other branching structures in nature, such as bronchial trees in the lungs or even root systems in botany?
This work is a significant step forward because it moves away from treating 3D shapes as simple point clouds and instead respects the underlying biological hierarchy of the subject matter. It is a clever, well-structured piece of engineering that sets a new standard for medical data synthesis.
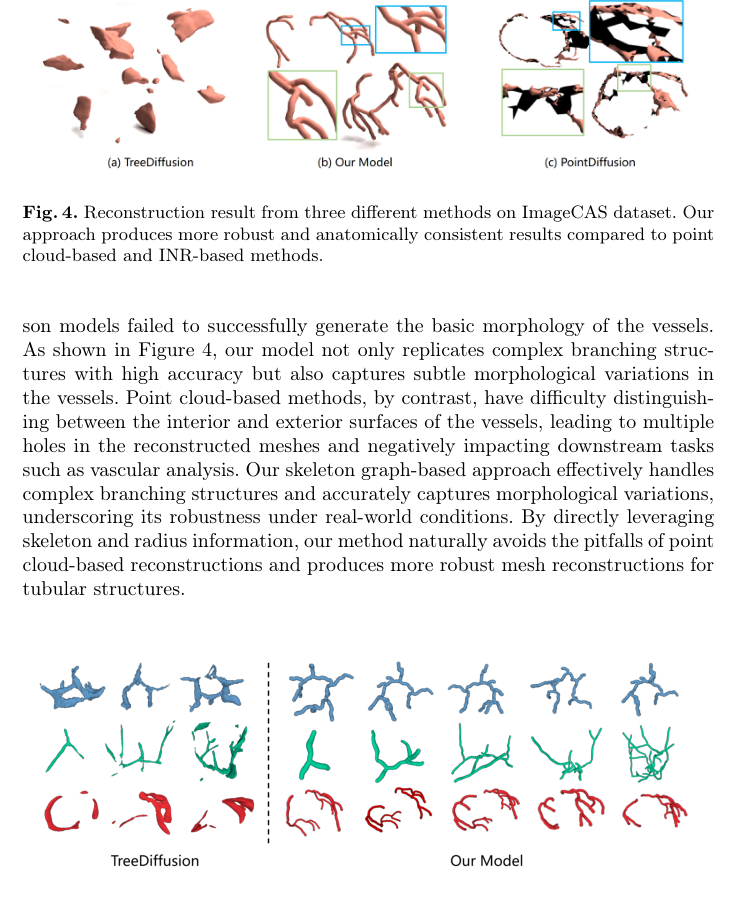 Figure 5. compares the generative performance of our approach against the highly competitive TreeDiffusion, using TreeDiffusion’s best-performing samples. As shown, TreeDiffusion often produces irregular, block-like shapes and dis- connected components across all datasets, indicating structural anomalies. In
Figure 5. compares the generative performance of our approach against the highly competitive TreeDiffusion, using TreeDiffusion’s best-performing samples. As shown, TreeDiffusion often produces irregular, block-like shapes and dis- connected components across all datasets, indicating structural anomalies. In
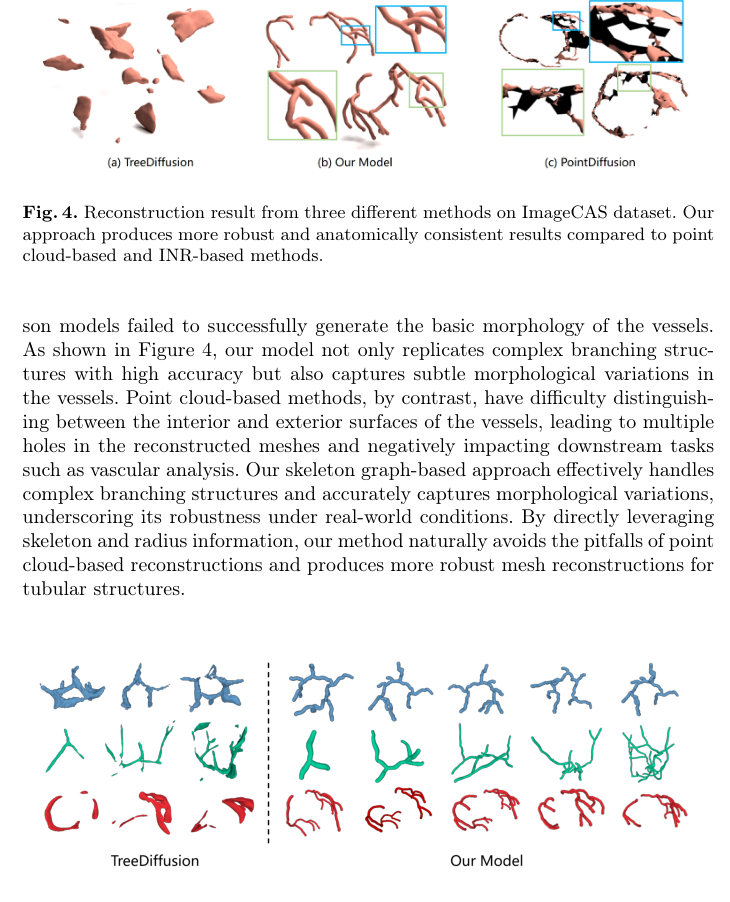 Figure 5. Examples of generation results from TreeDiffusion and our model on CoW, VascuSynth, and ImageCAS datasets (from top to bottom)
Figure 5. Examples of generation results from TreeDiffusion and our model on CoW, VascuSynth, and ImageCAS datasets (from top to bottom)
