RedDino: A foundation model for red blood cell analysis
RedDino analyzes red blood cell images with unprecedented accuracy, paving the way for faster disease diagnosis.
Background & Academic Lineage
The problem of automated red blood cell (RBC) analysis emerged from the clinical necessity to standardize hematological diagnostics, which are traditionally prone to human error and high variability. Historically, blood smear analysis relied on manual microscopic examination, a process heavily influenced by staining protocols and physical slide preparation. These factors introduce "batch effects"—variations in image appearance that make it difficult for AI models to generalize across different laboratories or patients. While foundation models have revolutionized general computer vision, their application to RBCs remained limited because existing models were not specifically tuned to the subtle, nuanced morphological differences required to distinguish between healthy and pathological cells.
The primary "pain point" addressed by the authors is that previous models often failed to generalize across diverse data sources due to these batch effects and the lack of a specialized, large-scale foundation model for RBCs. Furthermore, standard regularization techniques used in natural image models (like the Koleo regularizer) were actually counterproductive here, as they forced a uniformity that suppressed the very morphological abnormalities (e.g., malaria-infected cells or echinocytes) that clinicians need to detect.
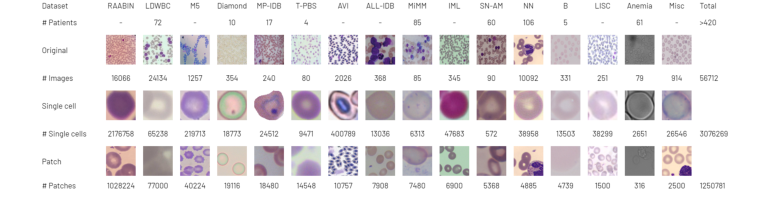 Figure 1. The RedDino training set comprises 56,712 original images. We extracted over 3 million single RBC images and more than 1.25 million patches
Figure 1. The RedDino training set comprises 56,712 original images. We extracted over 3 million single RBC images and more than 1.25 million patches
Intuitive Domain Terms
- Foundation Model: Think of this as a "general-purpose expert" that has been trained on a massive amount of data to understand the fundamental building blocks of images, which can then be easily "fine-tuned" to become a specialist in a specific task like identifying blood diseases.
- Self-Supervised Learning: Imagine teaching a student by giving them a giant pile of unlabeled puzzles. Instead of a teacher telling them what each piece is, the student learns by figuring out how the pieces fit together on their own. This allows the model to learn features without needing expensive, human-labeled data.
- Batch Effect: This is like trying to compare photos taken with different cameras, under different lighting, and with different filters. The "effect" is the artificial difference in the images caused by the equipment or process, rather than the actual content of the blood cells, which confuses the AI.
- Linear Probing: This is a simple test to see how "smart" the model's internal representation is. You freeze the model's brain and only train a very simple, shallow layer on top of it. If this simple layer performs well, it proves the model has already learned high-quality, useful features.
Notation Table
| Variable/Parameter | Description |
|---|---|
| $wF1$ | Weighted F1-score, a metric balancing precision and recall for imbalanced classes. |
| $bAcc$ | Balanced accuracy, which accounts for class imbalance by averaging recall per class. |
| $Acc$ | Standard accuracy, the ratio of correct predictions to total samples. |
| $N$ | Number of images or patches used in training or testing. |
| $d$ | The feature dimension (e.g., 384, 768, or 1024) of the model's embedding space. |
The authors solved the problem of feature representation for RBCs by adapting the DINOv2 framework. The core challenge was to move away from standard natural image training, which relies on the Koleo regularizer to prevent "feature collapse" (where the model maps all inputs to the same point). The authors identified that for RBCs, this regularizer was detrimental.
They replaced the standard moving average centering with the Sinkhorn-Knopp algorithm. Mathematically, this involves solving an optimal transport problem to map the feature distribution to a uniform distribution over the batch, ensuring that the model maintains distinct clusters for different cell types rather than collapsing them. By removing the Koleo regularizer and implementing this specific centering, they allowed the model to preserve the subtle morphological variations necessary for clinical diagnosis.
The model architecture was scaled into three versions:
1. RedDino Small: $d=384$, 22 million parameters.
2. RedDino Base: $d=768$, 86 million parameters.
3. RedDino Large: $d=1024$, 304 million parameters.
By training on over 1.25 million patches, the authors created a robust backbone that outperforms existing state-of-the-art models by consistently achieving higher $wF1$ and $bAcc$ scores across multiple independent test datasets. This confirms that their approach of "patch-based" training, rather than individual cell segmentation, better captures the context needed for accurate hematological analysis.
Problem Definition & Constraints
Core Problem Formulation & The Dilemma
The primary objective of this research is to develop a robust, generalized "foundation model" for Red Blood Cell (RBC) analysis that can accurately classify cell morphology across diverse clinical settings. The starting point (Input) is a massive, heterogeneous collection of raw blood smear images sourced from various laboratories, imaging modalities, and staining protocols. The desired endpoint (Output) is a high-quality, universal feature representation (embedding) that remains invariant to the "batch effect"—the technical noise introduced by different microscopes or preparation methods—while remaining sensitive to subtle, clinically relevant morphological variations.
The fundamental dilemma lies in the trade-off between feature invariance and discriminative power. In standard self-supervised learning (like the original DINOv2), models are often trained with regularizers (e.g., the Koleo regularizer) designed to prevent "feature collapse" by forcing a uniform distribution of embeddings. However, the authors discovered that for RBCs, this uniformity is a liability. Because healthy RBCs are naturally uniform in shape and color, a model that enforces strict uniformity ends up suppressing the very features that define pathological or abnormal cells.
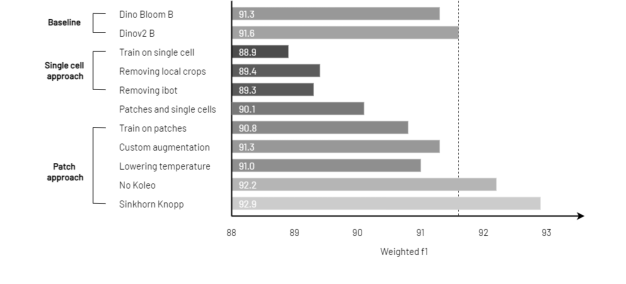 Figure 2. RedDino outperforms baseline models in linear probing evaluation by removing the Koleo regularizer and applying the Sinkhorn Knopp algorithm. The evaluation uses source 1 of the Elsafty dataset as the training set and source 2 as the test set
Figure 2. RedDino outperforms baseline models in linear probing evaluation by removing the Koleo regularizer and applying the Sinkhorn Knopp algorithm. The evaluation uses source 1 of the Elsafty dataset as the training set and source 2 as the test set
To bridge the gap between raw, noisy data and a robust feature space, the authors modified the DINOv2 framework through two critical interventions:
- Removal of the Koleo Regularizer: By removing the Koleo regularizer, the authors allowed the model to avoid the "over-suppression" of abnormal cell features. Mathematically, this allows the embedding space to retain higher variance, which is essential for distinguishing pathological cells that deviate from the "normal" cluster.
- Sinkhorn-Knopp Centering: The authors replaced the standard moving average centering with the Sinkhorn-Knopp algorithm. This is a classic approach for solving the Optimal Transport problem. In this context, it acts as a normalization technique that maps the feature distribution to a target distribution (often a uniform one) in a way that is more stable and better suited for the specific geometry of RBC data.
The authors also shifted the training strategy from individual cell crops to patched smear images. This allows the model to learn context—such as the relationship between cells and the background—which is crucial for distinguishing between true biological structures and artifacts.
Why This Approach
The development of RedDino represents a strategic pivot from general-purpose computer vision to a domain-specific foundation model for hematology. The authors identified that while models like DINOv2 are powerful, they are not inherently optimized for the specific morphological nuances of red blood cells (RBCs).
The Logic of the Approach
The authors determined that traditional "SOTA" methods were insufficient because they often rely on regularizers—specifically the Koleo regularizer—that are designed to prevent feature collapse in natural images by enforcing a uniform distribution of features. In the context of RBCs, this is counterproductive. Because RBCs exhibit a high degree of natural uniformity in shape and color, a regularizer that forces uniformity effectively suppresses the very features (pathological or abnormal variations) that are critical for clinical diagnosis. By removing this regularizer, the authors allowed the model to preserve the subtle, non-uniform morphological markers that distinguish healthy cells from diseased ones.
Comparative Superiority and Structural Advantages
- Sinkhorn-Knopp Centering: The authors replaced the standard moving average centering used in DINOv2 with the Sinkhorn-Knopp algorithm. This change provides a more robust way to handle the centering of feature distributions, which directly improves the quality of the learned representations in the presence of the high-dimensional noise typical of medical imaging.
- Custom Augmentation: By replacing standard pixel-level augmentations with a specialized pipeline of 32-pixel-level augmentations from the Albumentations library, the model becomes significantly more resilient to the artifacts introduced during the physical preparation of blood smears.
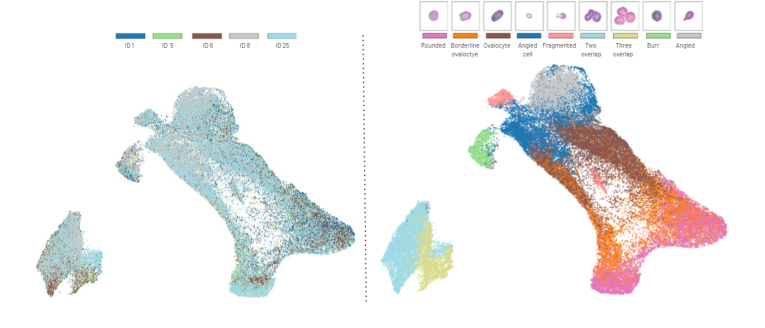 Figure 4. Different classes show distinct clusters in the UMAP projection of the feature embeddings from the Elsafty dataset source 1. On the left, we show the subject distri- bution across the UMAP space (each patient identified by a unique identifier), while on the right, we show the class distribution
Figure 4. Different classes show distinct clusters in the UMAP projection of the feature embeddings from the Elsafty dataset source 1. On the left, we show the subject distri- bution across the UMAP space (each patient identified by a unique identifier), while on the right, we show the class distribution
Mathematical & Logical Mechanism
The Mathematical Engine of RedDino
RedDino adapts the DINOv2 self-supervised learning framework to the domain of red blood cell (RBC) morphology. At its core, the model relies on a Self-Distillation with No Labels (DINO) mechanism, which functions as a teacher-student architecture.
The Master Equation
The objective function governing the training of the RedDino student network $g_{\theta_s}$ is to match the output distribution of a teacher network $g_{\theta_t}$. The core mechanism is the minimization of the cross-entropy loss between the teacher's soft probability distribution and the student's prediction:
$$ \mathcal{L} = - \sum_{x \in \{x_1, x_2\}} P_t(x) \log P_s(x) $$
Where:
1. $P_t(x) = \text{softmax}\left(\frac{f_{\theta_t}(x)}{T_t}\right)$ is the teacher's output distribution.
2. $P_s(x) = \text{softmax}\left(\frac{f_{\theta_s}(x)}{T_s}\right)$ is the student's output distribution.
Optimization Dynamics
The model learns by navigating a loss landscape defined by the similarity of RBC features. By removing the Koleo regularizer—which usually forces a uniform distribution of features—the authors allowed the model to preserve the natural clustering of RBCs. The Sinkhorn-Knopp algorithm replaces the standard moving average centering, acting as a constraint that forces the model to map inputs to a balanced distribution in the feature space, effectively preventing the model from "collapsing" into a single trivial solution.
Results, Limitations & Conclusion
The paper introduces RedDino, a specialized foundation model designed to address the challenges of red blood cell (RBC) morphological analysis.
Experimental Proof
The authors tested their model by comparing it against ResNet50, DINOv2 (the generic version), and DinoBloom (the current state-of-the-art for hematology). They used a "cross-source" evaluation strategy: they trained the model on one source of data and tested it on completely different, unseen sources. RedDino consistently outperformed the baselines, showing an average improvement of over 2% in linear probing and over 3% in K-NN metrics.
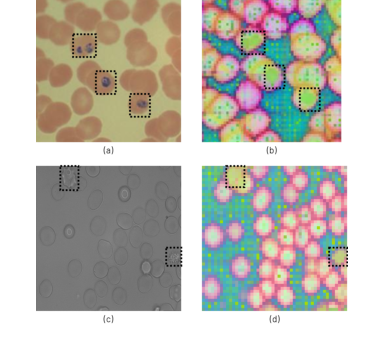 Figure 3. Abnormal RBC distinguished by RedDino: the highlighted regions in (a) and (c) correlate with distinct colors in the PCA visualization (b) and (d), showcasing their differentiation provided in the embedding space. Specifically, (a) contains malaria- infected RBCs, while (c) includes echinocytes
Figure 3. Abnormal RBC distinguished by RedDino: the highlighted regions in (a) and (c) correlate with distinct colors in the PCA visualization (b) and (d), showcasing their differentiation provided in the embedding space. Specifically, (a) contains malaria- infected RBCs, while (c) includes echinocytes
Discussion and Future Evolution
The success of RedDino opens several fascinating avenues for future research:
* Integration with Multimodal Data: Future iterations could integrate clinical metadata (e.g., patient age, hemoglobin levels) into the embedding space.
* Active Learning for Rare Pathologies: Since the model is already excellent at identifying outliers, it could be used in an active learning loop to automatically flag rare, undiagnosed blood conditions for human pathologists to review.
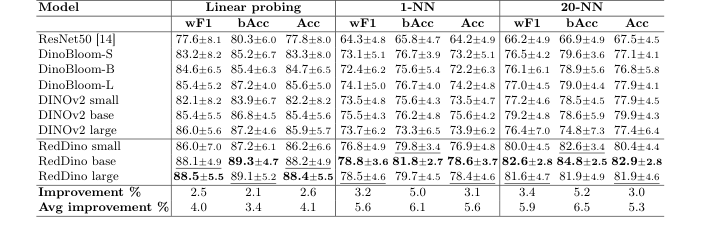 Table 1. RedDino models outperform ResNet50, DINOv2, and DinoBloom by over 2.1% in linear probing evaluation, and over 3.0% in 1-NN and 20-NN evaluation on the Elsafty dataset using a five-fold cross-validation strategy, where one source is fixed for training and the others are used for testing. "Avg Improvement" shows the average performance gain over baselines, while "Improvement" represents the performance gain compared to non-RedDino models
Table 1. RedDino models outperform ResNet50, DINOv2, and DinoBloom by over 2.1% in linear probing evaluation, and over 3.0% in 1-NN and 20-NN evaluation on the Elsafty dataset using a five-fold cross-validation strategy, where one source is fixed for training and the others are used for testing. "Avg Improvement" shows the average performance gain over baselines, while "Improvement" represents the performance gain compared to non-RedDino models
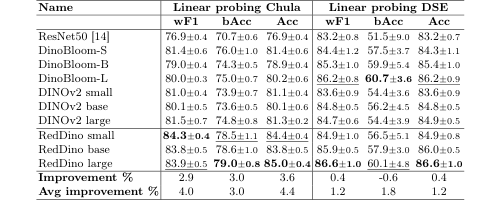 Table 2. RedDino outperforms baseline models in linear probing evaluations, with the only exception of the bAcc on the DSE dataset, in a five-fold cross-validation approach. "Avg Improvement" shows the average performance gain over baselines, while "Improvement" represents the performance gain compared to non-RedDino models
Table 2. RedDino outperforms baseline models in linear probing evaluations, with the only exception of the bAcc on the DSE dataset, in a five-fold cross-validation approach. "Avg Improvement" shows the average performance gain over baselines, while "Improvement" represents the performance gain compared to non-RedDino models
