MeDi: Metadata-Guided Diffusion Models for Mitigating Biases in Tumor Classification
Deep learning models have made significant advances in histological prediction tasks in recent years.
Background & Academic Lineage
The problem of "shortcut learning" in medical AI stems from the fact that deep learning models are often too efficient at finding patterns—so efficient that they latch onto irrelevant environmental noise rather than the actual biological signal. In histopathology, this means a model might learn to identify a tumor based on the specific color tint of a scanner or the staining protocol of a particular hospital, rather than the cellular morphology of the cancer itself. Historically, this emerged as AI moved from controlled laboratory datasets to real-world clinical deployment, where data heterogeneity is the norm. The fundamental "pain point" is that when training data is skewed (e.g., one hospital provides all the samples for a specific cancer type), the model creates a spurious correlation between the metadata (the hospital) and the target (the cancer). Consequently, when the model encounters a new hospital, it fails catastrophically because its "shortcut" no longer exists.
Intuitive Domain Terms
- Shortcut Learning: Imagine a student who passes a math exam not by learning algebra, but by memorizing that the answer to every question on page 1 is "5." The student performs perfectly on the practice test but fails the real exam because the questions are in a different order.
- Diffusion Model: Think of this as a sculptor working in reverse. You start with a block of random, noisy marble (random noise) and, guided by a set of instructions (metadata/class labels), you slowly chip away the noise until a clear, detailed statue (the medical image) remains.
- Subpopulation Shift: This is like training a chef to cook only with ingredients from a specific local market. If you suddenly move that chef to a different country with entirely different produce, they struggle to cook the same dish because they never learned to adapt to the new, unfamiliar ingredients.
Notation Table
| Notation | Description |
|---|---|
| $\alpha_k$ | The $k$-th metadata attribute (e.g., hospital site, patient race). |
| $d_e$ | The fixed dimension of the learnable embedding vector for categorical metadata. |
| $\mathbf{z}_{\text{site}(i)}$ | The embedding vector representing a specific medical center $i$. |
| $\mathbf{z}_{\text{class}}$ | The embedding vector representing the disease/cancer subtype. |
| $\mathbf{z}_{\text{meta},i}$ | The embedding vector for the $i$-th metadata attribute. |
| $\mathbf{z}_t$ | The timestep embedding vector used in the diffusion process. |
| $\mathbf{z}_{\text{cond}}$ | The final concatenated conditioning vector used to guide generation. |
| $\mathbf{z}_{\text{final}}$ | The combined vector $\mathbf{z}_t + \mathbf{z}_{\text{cond}}$ provided to the UNet blocks. |
The authors solve the bias problem by explicitly injecting metadata into the generative process. Instead of a standard diffusion model that only learns the mapping $p(\text{image} \mid \text{class})$, they redefine the objective to learn $p(\text{image} \mid \text{class}, \text{metadata})$.
They achieve this by creating a conditioning vector $\mathbf{z}_{\text{cond}}$ that merges the class information with all relevant metadata attributes:
$$\mathbf{z}_{\text{cond}} = \text{concat}(\mathbf{z}_{\text{class}}, \mathbf{z}_{\text{meta},1}, \dots, \mathbf{z}_{\text{meta},k}) \in \mathbb{R}^{d_t}$$
This vector is then integrated into the UNet's internal denoising process by adding it to the timestep embedding $\mathbf{z}_t$:
$$\mathbf{z}_{\text{final}} = \mathbf{z}_t + \mathbf{z}_{\text{cond}}$$
By doing this, the model is forced to learn how specific metadata (like a hospital's unique staining style) interacts with the biological features of the tissue. During inference, the user can "mix and match" these conditions to generate synthetic data for underrepresented or unseen combinations, effectively balancing the dataset and forcing the downstream classifier to ignore the metadata shortcuts.
Problem Definition & Constraints
Core Problem Formulation & The Dilemma
The Starting Point (Input/Current State):
In clinical histopathology, deep learning models are trained on large datasets (like TCGA) to perform diagnostic tasks, such as tumor subtyping. These datasets are inherently skewed, as they aggregate data from various medical centers, each with unique staining protocols, scanner hardware, and patient demographics.
The Desired Endpoint (Output/Goal State):
The goal is to create a robust diagnostic model that generalizes across diverse clinical environments. Specifically, the authors aim to generate high-fidelity synthetic histopathology images that represent underrepresented or entirely unseen subpopulations (e.g., a specific cancer type from a hospital not present in the training set). By augmenting the training data with these synthetic samples, the model should achieve a balanced distribution, effectively "filling in" the gaps in the data.
The Missing Link:
The gap lies in the inability of standard generative models to decouple biological features (the disease) from metadata-driven variations (the "domain" or "site" effects). When a model is trained on a skewed dataset, it fails to distinguish between the actual tumor morphology and the spurious correlations introduced by the specific site's imaging artifacts.
The Dilemma (The Trade-off):
Researchers face a classic "shortcut learning" trap. If a model is trained to classify tumors, it often learns to rely on the metadata (e.g., "this specific staining pattern belongs to Hospital A") as a proxy for the label. If you force the model to ignore these variations, you lose the ability to generate realistic, site-specific images. Conversely, if you allow the model to learn these variations, it becomes biased and fails to generalize to new, unseen hospitals.
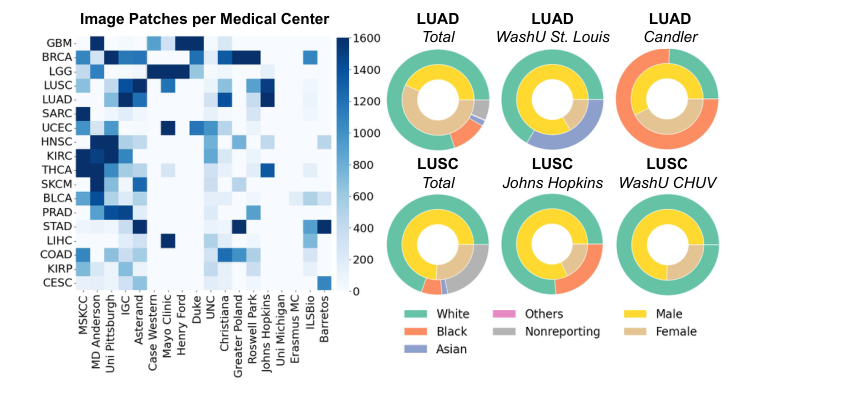 Figure 2. Dataset statistics for TCGA-UT. Left: The number of patches for the 18 largest cancer types (vertical axis) and tissue source sites (horizontal axis), capped at 1,600. Right: Race and gender distributions for the patches of selected cancer types and tissue source sites. The dataset is highly imbalanced across classes, hospitals, and demographics, with many missing or underrepresented metadata combinations
Figure 2. Dataset statistics for TCGA-UT. Left: The number of patches for the 18 largest cancer types (vertical axis) and tissue source sites (horizontal axis), capped at 1,600. Right: Race and gender distributions for the patches of selected cancer types and tissue source sites. The dataset is highly imbalanced across classes, hospitals, and demographics, with many missing or underrepresented metadata combinations
The Harsh, Realistic Walls:
1. Combinatorial Explosion: The metadata space is massive. With 626 tissue source sites and 32 cancer types, the potential combinations ($626 \times 32 = 20,032$) are only partially represented in real-world data. This makes it impossible to rely on simple data collection to cover all scenarios.
2. Spurious Correlations: The data is highly imbalanced; certain cancer types are exclusively associated with specific hospitals in the training set. This creates a "Clever Hans" effect where the model learns to associate the hospital's unique "look" with the cancer type, rather than the biological features of the cancer itself.
3. Non-Differentiable/Discrete Metadata: Integrating categorical metadata (like hospital IDs) into a continuous diffusion process requires a carefully designed embedding strategy to ensure the model can effectively condition the generation process without collapsing into a single "mode" of the data distribution.
Why This Approach
The core challenge addressed by this paper is the "Clever Hans" effect in computational pathology, where deep learning models inadvertently learn to rely on non-biological metadata—such as hospital-specific staining protocols, scanner artifacts, or demographic skews—rather than the actual tumor morphology. When a model is trained on a dataset where specific cancer types are correlated with specific hospitals, it treats these metadata as shortcuts, leading to catastrophic failure when deployed in a new clinical environment with a different data distribution.
The Inevitability of the Choice
The authors identified that standard "SOTA" approaches, including large-scale foundation models trained via self-supervised learning, are insufficient because they implicitly encode these metadata biases into their latent representations. If the training distribution is skewed, these models simply inherit the bias. The authors realized that to truly mitigate this, one cannot rely on passive learning; one must explicitly model the metadata as a conditioning variable.
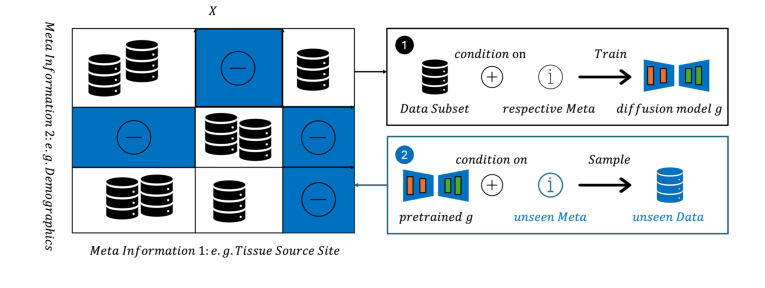 Figure 1. MeDi Training and Inference Framework. During training (1), the model receives real images along with their class labels and metadata. At inference (2), users can condition on arbitrary combinations of class and metadata, enabling the genera- tion of synthetic images for underrepresented subpopulations or unseen subpopulations altogether
Figure 1. MeDi Training and Inference Framework. During training (1), the model receives real images along with their class labels and metadata. At inference (2), users can condition on arbitrary combinations of class and metadata, enabling the genera- tion of synthetic images for underrepresented subpopulations or unseen subpopulations altogether
- Comparative Superiority: Unlike traditional stain normalization or style-transfer techniques (e.g., CycleGAN), which attempt to "fix" the image by forcing it into a canonical style, MeDi treats metadata as a controllable parameter. By using a diffusion model conditioned on both class labels and metadata (e.g., Tissue Source Site), the framework gains the ability to perform targeted data augmentation. It can interpolate within the metadata space to balance existing combinations or extrapolate to generate synthetic samples for underrepresented or entirely unseen subpopulations. This structural advantage allows the model to "fill in" the gaps in the training distribution, effectively decoupling the disease label from the hospital-specific artifacts.
- The "Marriage" of Requirements and Solution: The problem requires a generative model that is both high-fidelity and highly controllable. Diffusion models are the only viable solution here because they provide a stable, iterative denoising process that can be easily conditioned at every step. By defining a conditioning vector $\mathbf{z}_{\text{cond}} = \text{concat}(\mathbf{z}_{\text{class}}, \mathbf{z}_{\text{meta},1}, \dots, \mathbf{z}_{\text{meta},k})$ and injecting it into the UNet's residual blocks via $\mathbf{z}_{\text{final}} = \mathbf{z}_t + \mathbf{z}_{\text{cond}}$, the authors ensure that the generation process is strictly guided by the desired metadata. This perfectly aligns with the need to synthesize images that maintain biological integrity while exhibiting the specific "style" of an underrepresented hospital site.
Mathematical & Logical Mechanism
To understand this paper, one must first grasp the concept of "shortcut learning" in medical AI. When a model is trained to classify tumors, it often inadvertently learns to associate specific hospital-related artifacts (like staining colors or scanner noise) with a disease label, rather than learning the actual biological features of the cancer. This happens because certain hospitals might only submit specific types of cancer, creating a false correlation. The authors propose MeDi to break these correlations by explicitly injecting metadata (like the hospital site) into the generative process, allowing the model to "decouple" the disease from the site-specific noise.
The Master Equation
The core of the MeDi framework is the construction of a conditioning vector that guides the diffusion model's denoising process. The final conditioning signal provided to the UNet is defined as:
$$ \mathbf{z}_{\text{final}} = \mathbf{z}_t + \mathbf{z}_{\text{cond}} $$
Where $\mathbf{z}_{\text{cond}}$ is defined as:
$$ \mathbf{z}_{\text{cond}} = \text{concat}(\mathbf{z}_{\text{class}}, \mathbf{z}_{\text{meta},1}, \dots, \mathbf{z}_{\text{meta},k}) \in \mathbb{R}^{d_t} $$
Tearing the Equation Apart
- $\mathbf{z}_t$: This is the timestep embedding. It represents the current "noise level" in the diffusion process. Its role is to inform the model how much denoising is required at the current step.
- $\mathbf{z}_{\text{class}}$: This is the learnable embedding of the cancer subtype (e.g., lung adenocarcinoma). It provides the primary semantic guidance for what biological structure to generate.
- $\mathbf{z}_{\text{meta},i}$: These are the learnable embeddings for the $k$ metadata attributes (e.g., tissue source site). Their role is to act as a "style" or "domain" controller, forcing the model to learn the specific visual artifacts associated with a particular hospital.
- $\text{concat}(\dots)$: The author uses concatenation to fuse these distinct information sources into a single vector. This is preferred over addition here because the class and metadata represent independent, categorical dimensions that should not be mixed until the model explicitly processes them within the UNet layers.
- $\mathbf{z}_{\text{final}}$: This is the combined conditioning vector. By adding it to $\mathbf{z}_t$, the authors ensure that the denoising operation is simultaneously aware of the "time" (noise level) and the "context" (class + metadata).
Results, Limitations & Conclusion
In computational pathology, deep learning models often suffer from "shortcut learning." Because medical datasets are typically collected from specific hospitals, they contain inherent biases—such as unique staining protocols, scanner artifacts, or demographic skews—that correlate with disease labels. A model might learn to identify a tumor not by its biological morphology, but by the specific "look" of the tissue slides from a particular hospital. When deployed in a new environment, these models fail because they rely on these spurious correlations rather than the underlying pathology.
Experimental Validation
The authors "ruthlessly" tested their hypothesis by creating a challenging out-of-distribution scenario. They held out 30% of specific medical center and patient race combinations, ensuring the model had never seen these specific subpopulations during training.
- The Evidence:
- Fidelity: MeDi achieved a lower average Fréchet Inception Distance (FID) of 37.73 compared to the CLS baseline's 50.65, proving that metadata conditioning leads to more faithful image synthesis.
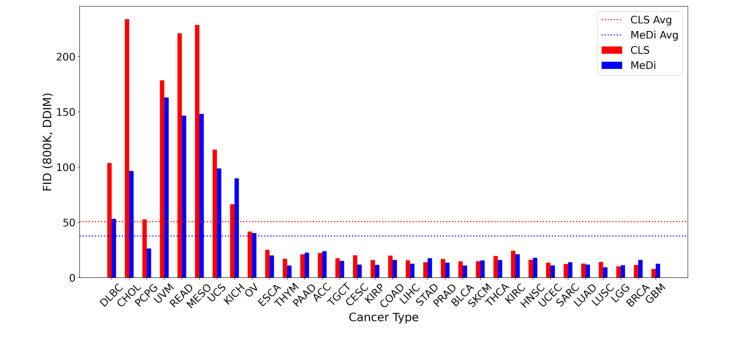 Figure 3. FID per cancer type at 800K optimization steps sampled with DDIM. Depicted are the class-only baseline (CLS, red) and the metadata-conditioned model (MeDi, blue). Cancer types are ordered in descending order based on the number of images in the dataset. Dotted horizontal lines represent the average FID per model: CLS: 50.65, MeDi: 37.73
Figure 3. FID per cancer type at 800K optimization steps sampled with DDIM. Depicted are the class-only baseline (CLS, red) and the metadata-conditioned model (MeDi, blue). Cancer types are ordered in descending order based on the number of images in the dataset. Dotted horizontal lines represent the average FID per model: CLS: 50.65, MeDi: 37.73
2. **Downstream Utility:** The authors trained linear classifiers on top of embeddings from a foundation model (UNI). When tested on unseen subpopulations, the MeDi-augmented training set consistently outperformed the CLS-augmented set in balanced accuracy for NSCLC and Uterine cancer tasks. This provides definitive proof that MeDi successfully breaks the spurious correlations that usually plague these models.
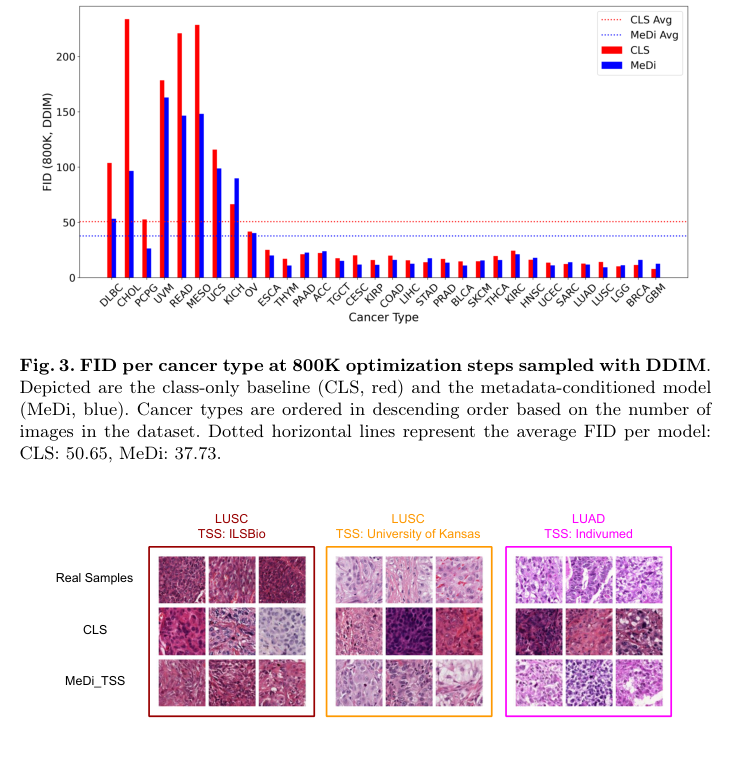 Figure 4. LUSC and LUAD samples from different tissue source sites (TSS) along with generated images. The top row shows real data, while the two other rows show generated samples. MeDi enables capturing the staining mode of the different tissue source sites, while the CLS only model can not be conditioned on a specific TSS and, therefore, is unable to match the real data sample distribution
Figure 4. LUSC and LUAD samples from different tissue source sites (TSS) along with generated images. The top row shows real data, while the two other rows show generated samples. MeDi enables capturing the staining mode of the different tissue source sites, while the CLS only model can not be conditioned on a specific TSS and, therefore, is unable to match the real data sample distribution
The authors effectively demonstrated that by explicitly modeling the "noise" (metadata), they could force the model to focus on the "signal" (pathology), resulting in a more robust and fair system.
