Hybrid Graph Mamba: Unlocking Non-Euclidean Potential for Accurate Polyp Segmentation
Colorectal polyp segmentation can assist doctors in screening colonoscopy images, which is crucial for the prevention of colorectal cancer.
Background & Academic Lineage
Colorectal polyp segmentation is a critical task in medical imaging, primarily aimed at assisting doctors in screening colonoscopy images. The ultimate goal is to prevent colorectal cancer (CRC), which is a leading cause of cancer-related deaths globally. Polyps are small growths in the colon that can develop into cancer, so their early and accurate detection is paramount.
Historically, the process of identifying and outlining polyps in colonoscopy images was performed manually by doctors. This approach, while standard, suffered from several significant drawbacks. It was incredibly time-consuming, labor-intensive, and highly subjective, meaning that different doctors might interpret the same image differently. This subjectivity and the sheer volume of images often led to missed detections, which could have severe consequences for patients.
To overcome these human limitations, automated polyp segmentation techniques emerged. Early attempts relied on "handcrafted features," where engineers would manually design algorithms to recognize specific patterns, shapes, or textures associated with polyps. However, these methods were quite limited in their ability to capture the complex and varied appearance of polyps, often leading to high rates of false positives or missed polyps.
The advent of deep learning brought a significant leap forward, allowing models to learn more comprehensive and accurate features directly from data. Despite these advancements, several fundamental limitations, or "pain points," persisted, which this paper directly addresses:
- Neglect of Non-Euclidean Features: Most existing deep learning methods primarily focused on "Euclidean features," like the simple shape, size, and texture of a polyp. However, they largely ignored "non-Euclidean features," which describe the more complex geometric and topological relationships between a polyp and its surrounding tissue. Imagine not just the polyp itself, but how it's connected to the colon wall, its irregular surface, or how it interacts with nearby folds. These relational features are crucial for accurate segmentation but were often overlooked.
- Ineffective Feature Fusion for Non-Euclidean Structures: Non-Euclidean features are not uniform; they vary significantly across different regions of an image (e.g., the polyp's interior, its edges, or the background). Previous methods for combining features often treated all regions uniformly, failing to account for these regional differences and the unique topological structures present in non-Euclidean data. This meant valuable contextual information was lost or poorly integrated.
- Underutilization of Low-Level Features and Bridging the Feature Gap: Deep learning models typically extract features at multiple levels: "low-level" features capture fine details like edges and textures, while "high-level" features capture broader semantic information (e.g., "this is definitely a polyp"). Existing methods often didn't fully exploit these low-level details or struggled to effectively combine them with high-level semantic information, leading to less precise boundary delineation and overall segmentation.
These three issues collectively represent the core motivation for the authors to develop their Hybrid Graph Mamba (HGM) model, aiming to unlock the potential of non-Euclidean features for more accurate polyp segmentation.
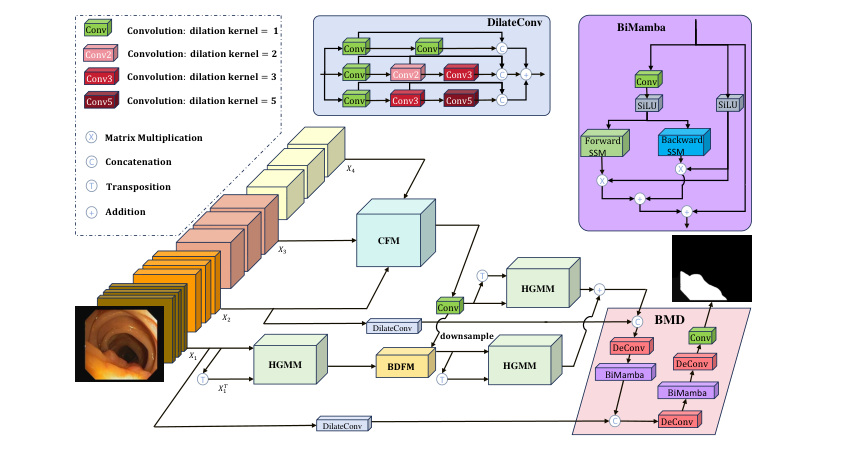 Figure 1. Overall architecture of HGM. Our model consists of a pyramid vision trans- former, a CFM, three HGMMs, a BDFM, and a BMD
Figure 1. Overall architecture of HGM. Our model consists of a pyramid vision trans- former, a CFM, three HGMMs, a BDFM, and a BMD
Problem Definition & Constraints
Core Problem Formulation & The Dilemma
The Starting Point (Input): The model receives raw colonoscopy images containing polyps of varying sizes, shapes, and textures, often obscured by ambiguous boundaries against surrounding colonic tissue.
The Desired Endpoint (Output): A precise binary segmentation mask that accurately delineates the polyp region from the background, capturing both global semantic context and fine-grained boundary details.
The Missing Link: The fundamental gap lies in the inability of traditional deep learning architectures to simultaneously capture Euclidean features (local texture and shape) and non-Euclidean features (the complex topological and geometric relationships between the polyp and its surrounding tissue). While standard CNNs or Transformers excel at local or global patterns, they often fail to model the irregular, graph-like structural dependencies inherent in biological tissues.
The Dilemma (The Trade-off): Researchers face a "representation bottleneck." Increasing the model's capacity to capture global semantic information (high-level features) typically leads to the loss of spatial resolution and boundary precision (low-level features). Conversely, focusing purely on high-resolution details often results in a lack of global context, causing the model to misidentify background noise as polyps.
The Harsh Constraints:
1. Topological Complexity: Polyps do not follow simple grid-like structures; their boundaries are highly irregular, making standard convolution kernels insufficient for capturing the "non-Euclidean" geometric relationships.
2. Feature Uniformity: Most existing fusion methods treat all feature levels (low-level details vs. high-level semantics) with the same mathematical weight, failing to account for the distinct roles of internal, edge, and background regions.
3. Computational Efficiency: Implementing graph-based operations or complex attention mechanisms often introduces prohibitive memory overhead, making it difficult to maintain real-time or near-real-time performance required for clinical settings. The authors had to design a sparse adjacency matrix to keep the GCN computation tractable, as a fully connected graph would be computationally infeasible for high-resolution medical images.
The authors bridge this gap by proposing the Hybrid Graph Mamba (HGM), which integrates Mamba (a state-space model) with Graph Convolutional Networks (GCN).
-
Quad-directional Mamba (QM): To address the limitation of standard sequential processing, the authors use a quad-directional approach to extract features from four directions. This allows the model to capture long-range dependencies in the image while maintaining a linear complexity, unlike the quadratic complexity of standard Transformers. The core operation is defined as:
$$ \text{BiMamba}(x) = \text{RS}(x + x' \text{SSM}_F(x'') + x' \text{SSM}_B(x'')) $$
where $x'$ and $x''$ are non-linear transformations of the input, and $\text{SSM}_F$ and $\text{SSM}_B$ represent the forward and backward state-space models. -
Non-Euclidean Feature Extraction: To explicitly model the non-Euclidean topology, the authors feed the concatenated directional features into a GCN. The output of the Hybrid Graph Mamba Module (HGMM) is defined as:
$$\text{HGMM}(X) = \text{GCN}([X_F, X_B, X_F^\top, X_B^\top], A) + X_M + X_M^\top + X$$
Here, $A$ is a sparse adjacency matrix where only specific positions (every 32 units) are set to one to reduce the computational burden while still capturing structural relationships. -
Boundary Discrimination Fusion (BDFM): To solve the fusion dilemma, the authors process high-level features to generate an initial segmentation map, which is then used to derive distinct feature maps for the internal, edge, and background regions. These are flattened into a tensor $U$ and fused with the low-level features $X'$ via a series of convolutions:
$$X_{\text{BDFM}} = \text{Conv}([\text{RS}(\text{Conv}(UX'))(\text{Conv}(UX')\text{Conv}(X')), \text{RS}(X')])$$
Why This Approach
The authors realized that existing SOTA methods, primarily based on standard Convolutional Neural Networks (CNNs) and even some early Transformer-based approaches, were insufficient due to three key limitations:
- Neglect of Non-Euclidean Features: Most methods focused solely on "Euclidean features" like the shape and texture of polyps. However, the geometric and topological relationships between a polyp and its surrounding tissue – which are "non-Euclidean features" – were largely ignored.
- Ineffective Feature Fusion for Regional Differences: Non-Euclidean features are not uniform; they vary significantly across different regions of an image (e.g., the polyp's interior, its edges, and the background). Existing feature fusion techniques often treated all features uniformly, failing to account for these vital regional distinctions.
- Underutilization of Low-Level Features and Gap Between Levels: Traditional methods often didn't fully exploit these low-level details or struggled to effectively bridge the information gap between low- and high-level features during fusion, leading to blurry boundaries or missed small polyps.
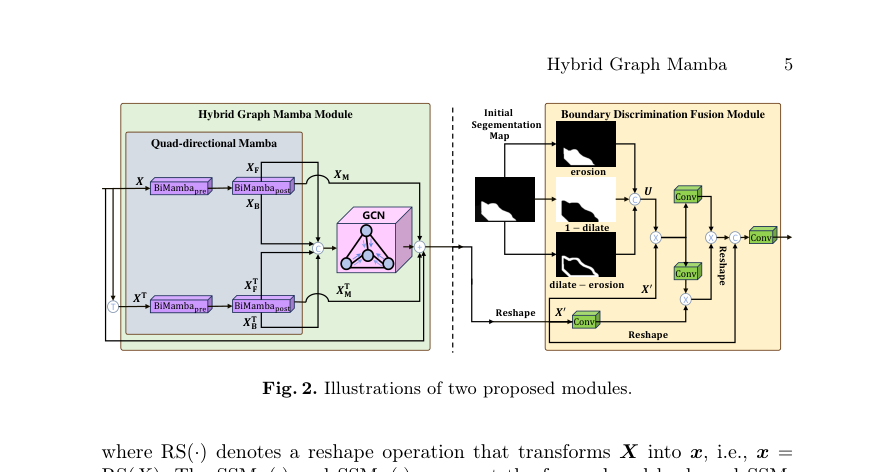 Figure 2. Illustrations of two proposed modules
Figure 2. Illustrations of two proposed modules
Comparative Superiority (The Benchmarking Logic)
The Hybrid Graph Mamba (HGM) method offers qualitative superiority by directly addressing the identified shortcomings:
- Explicit Non-Euclidean Feature Extraction: Unlike standard CNNs that operate on grid-like (Euclidean) data, HGM incorporates Graph Convolutional Networks (GCNs) within its Hybrid Graph Mamba Module (HGMM). GCNs are specifically designed to process graph-structured data, allowing HGM to explicitly model and extract the non-Euclidean geometric and topological relationships.
- Region-Aware Multi-Scale Fusion: HGM introduces a Boundary Discrimination Fusion Module (BDFM) that doesn't treat all features uniformly. Instead, it processes an initial segmentation map to derive distinct feature maps for the internal, edge, and background regions.
- Efficient Multi-Scale Feature Aggregation with Mamba: The integration of Mamba (specifically BiMamba blocks) provides a powerful mechanism for sequence modeling. Mamba's State Space Model (SSM) architecture offers linear complexity with respect to sequence length, a significant advantage over the quadratic complexity of self-attention in standard Transformers.
Mathematical & Logical Mechanism
The Hybrid Graph Mamba (HGM) model addresses the limitations of traditional deep learning in medical imaging by explicitly modeling non-Euclidean topological structures alongside standard Euclidean features.
The Master Equation
The core logic of the Hybrid Graph Mamba Module (HGMM), which serves as the primary engine for non-Euclidean feature extraction, is defined as:
$$\text{HGMM}(\mathbf{X}) = \text{GCN}([\mathbf{X}_F, \mathbf{X}_B, \mathbf{X}_F^\top, \mathbf{X}_B^\top], \mathbf{A}) + \mathbf{X}_M + \mathbf{X}_M^\top + \mathbf{X}$$
Tearing the Equation Apart
- $[\mathbf{X}_F, \mathbf{X}_B, \mathbf{X}_F^\top, \mathbf{X}_B^\top]$: This is the concatenation of four directional feature maps.
- $\mathbf{A}$: The adjacency matrix. It defines the graph structure. The authors set specific values to 1 (every 32 units) to enforce a sparse but meaningful connectivity.
- $\text{GCN}(\cdot, \mathbf{A})$: This operator performs graph convolution, aggregating information from neighboring nodes defined by $\mathbf{A}$.
- $\mathbf{X}_M + \mathbf{X}_M^\top$: These are the residual outputs from the post-BiMamba blocks, preserving sequential information.
- $+\mathbf{X}$: This is the final residual connection, preventing the vanishing gradient problem.
Results, Limitations & Conclusion
The authors "ruthlessly" tested their architecture against eight state-of-the-art (SOTA) models across five benchmark datasets (CVC-300, ClinicDB, Kvasir, ColonDB, and ETIS).
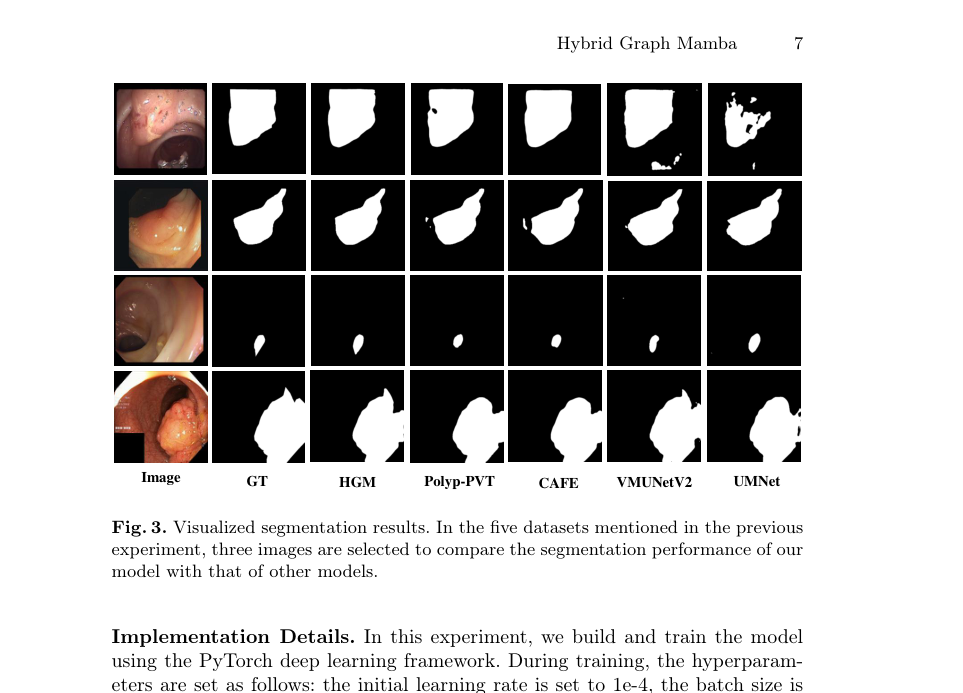 Figure 3. Visualized segmentation results. In the five datasets mentioned in the previous experiment, three images are selected to compare the segmentation performance of our model with that of other models
Figure 3. Visualized segmentation results. In the five datasets mentioned in the previous experiment, three images are selected to compare the segmentation performance of our model with that of other models
The definitive evidence of HGM's superiority is found in its consistent performance across all datasets. HGM achieves the best overall average (Dice: 0.887, IoU: 0.825). The ablation study in Table 2 serves as the "smoking gun," proving that adding each component—BMD, QM, GCN, and BDFM—incrementally improves the Dice and IoU metrics, confirming that the architectural choices are not just coincidental but mathematically sound.
