D3M: Deformation-Driven Diffusion Model for Synthesis of Contrast-Enhanced MRI with Brain Tumors
Contrast-enhanced magnetic resonance images (CEMRIs) provide valuable information for brain tumor diagnosis and treatment planning.
Background & Academic Lineage
The Origin & Academic Lineage
The core problem addressed in this paper is the synthesis of Contrast-Enhanced Magnetic Resonance Images (CEMRIs) from Non-Contrast Magnetic Resonance Images (NCMRIs), particularly for brain tumor diagnosis. This specific challenge emerged from a critical need in clinical practice: while CEMRIs provide invaluable information for identifying and planning treatment for brain tumors, their acquisition necessitates the injection of contrast agents. This procedure carries several significant drawbacks, including potential patient health risks [15, 25], substantial high costs [27], and growing environmental concerns related to the disposal of these agents [2, 8].
Historically, researchers have sought to circumvent these issues by developing methods to computationally generate CEMRIs from standard NCMRIs, thereby eliminating the need for contrast agent injection while preserving diagnostic image quality. Early efforts in this field leveraged deep learning architectures such as 3D Convolutional Neural Networks (CNNs) based on U-Net and conditional Generative Adversarial Networks (GANs), inspired by models like Pix2Pix [17, 6]. More recently, the academic lineage has seen a significant shift towards diffusion models, which have demonstrated remarkable success in generating realistic medical images [16, 24, 26, 29]. These state-of-the-art diffusion models have been adapted for CEMRI synthesis, with some approaches focusing on specific organs like the liver [26].
A fundamental limitation, or "pain point," of these previous approaches, including existing diffusion models, is that CEMRI synthesis from NCMRIs remains a highly ill-posed problem. NCMRIs often provide only ambiguous evidence regarding enhanced regions, making it difficult for models to accurately predict where contrast enhancement should occur. Consequently, prior methods frequently produce noticeable false positive and false negative enhancement results. This means they might incorrectly show enhancement in non-enhanced areas (false positives) or fail to show enhancement in areas that should be enhanced (false negatives). This issue is particularly pronounced in tumor regions, where the intricate and often complex morphology of tumor subcomponents is not accurately captured. The authors of this paper address this by reformulating the problem: instead of treating enhancement errors purely as intensity discrepancies (which are often large and hard to correct), they conceptualize them as incorrect geometric interpretations of tumor subcomponents. This allows for correction through spatial deformation, a more manageable task as the required geometric adjustments are typically smaller.
Intuitive Domain Terms
To help a zero-base reader grasp the concepts, here are some specialized terms translated into everyday analogies:
- Contrast-Enhanced Magnetic Resonance Images (CEMRIs): Imagine you're looking at a map of a city, but some important buildings (like hospitals or landmarks) are hard to distinguish. A CEMRI is like using a special highlighter to make those important buildings glow brightly, so they immediately stand out and are easy to identify.
- Non-Contrast Magnetic Resonance Images (NCMRIs): Following the map analogy, an NCMRI is just the regular map without any special highlighting. All the buildings are there, but the important ones don't stand out on their own.
- Diffusion Model: Think of a blurry, pixelated photo that slowly becomes clear and detailed as if an artist is meticulously adding details back in. A diffusion model is an AI that learns to do this in reverse: it starts with pure static (like a TV with no signal) and gradually "un-blurs" or "denoises" it step-by-step until it reveals a complete, realistic image.
- Ill-posed problem: This is like trying to solve a riddle where there are many possible answers, or where a tiny change in your initial guess leads to a completely different outcome. For CEMRI synthesis, it means that without the contrast agent, the input (NCMRI) doesn't give enough clear clues to reliably produce a perfect CEMRI, making it prone to errors.
- Spatial Deformation: Picture drawing a face on a balloon. Spatial deformation is like gently squeezing or stretching the balloon to subtly shift the position of the eyes or mouth, rather than erasing and redrawing them. It's about moving parts of an image around geometrically to correct their placement.
Notation Table
| Notation | Description |
|---|---|
Problem Definition & Constraints
Core Problem Formulation & The Dilemma
The core problem addressed by this paper is the synthesis of Contrast-Enhanced Magnetic Resonance Images (CEMRIs) from Non-Contrast Magnetic Resonance Images (NCMRIs), specifically T1-weighted, T2-weighted, and FLAIR images.
The starting point (input) for the model is a set of NCMRIs. During training, auxiliary information in the form of annotated masks of enhanced tumors is also utilized to guide the learning process. The desired endpoint (output) is a high-quality synthetic CEMRI that accurately depicts brain tumors and their enhancement patterns, without the need for actual contrast agent injection.
The exact missing link or mathematical gap lies in the inherent "ill-posed" nature of synthesizing CEMRIs from NCMRIs. NCMRIs provide only "ambiguous evidence about the enhanced regions," making it extremely challenging to accurately predict where and how contrast enhancement should appear. Previous research, including methods based on diffusion models, has struggled with this, frequently producing "false positive and false negative enhancement results." This means that non-enhanced regions might be incorrectly depicted with high intensities (false positives), while regions that should be enhanced are shown with low intensities (false negatives). The critical gap is the inability of these models to precisely capture the "intricate morphology of tumor subcomponents" and their specific enhancement characteristics.
The painful trade-off or dilemma that has trapped previous researchers stems from their approach to correcting these errors. They typically model enhancement discrepancies as intensity errors. However, correcting these intensity errors is "typically large and challenging to correct." This creates a difficult balancing act: improving the fidelity of enhancement often requires drastic and hard-to-manage intensity adjustments, which can lead to artifacts or an overall less realistic image. This paper proposes a novel reformulation: instead of viewing these as intensity errors, they are framed as an "incorrect interpretation of tumor subcomponents." This allows for geometric correction via spatial deformation, which is presented as "relatively small and more manageable" compared to large intensity corrections.
Constraints & Failure Modes
The problem of synthesizing CEMRIs from NCMRIs is made insanely difficult by several harsh, realistic constraints:
- Physical Constraints:
- Health Risks & Costs of Contrast Agents: The primary motivation for this work is to circumvent the need for gadolinium-based contrast agents, which carry "health risks, high costs, and environmental concerns." This constraint drives the need for accurate synthesis.
- Ambiguous Input Information: NCMRIs inherently provide "ambiguous evidence about the enhanced regions." This means the input data itself lacks the explicit information needed to directly infer contrast enhancement, making the synthesis task fundamentally challenging.
- Computational Constraints:
- Ill-Posed Problem: The synthesis of CEMRIs from NCMRIs is a "highly ill-posed" problem. This implies that multiple possible CEMRIs could correspond to a single NCMRIs input, making it difficult for a model to uniquely determine the correct enhanced image without additional guidance.
- Intricate Morphology: Existing models fail to capture the "intricate morphology of tumor subcomponents." This suggests that simple pixel-wise or intensity-based mappings are insufficient to represent the complex shapes and boundaries of enhanced tumor regions.
- Stepwise Denoising Process: Diffusion models operate through a "stepwise" denoising process. Integrating geometric correction effectively within these multiple steps without accumulating errors or disrupting the diffusion process is a significant computational and architectural challenge.
- Data-Driven Constraints:
- Need for High-Quality Annotations (for training): While the goal is to avoid contrast agents in inference, the model relies on "auxiliary information of annotations of enhanced tumors" during training. These masks must be "manually annotated and reviewed by clinical experts to ensure high-quality labels," which is a labor-intensive and costly process. The quality and availability of such precise ground truth data are critical for successful model training.
- Difficulty in Generalization: The complex and varied nature of brain tumors means that models must be robust enough to synthesize accurate enhancements across diverse tumor types and patient anatomies, which is a significant generalization challenge.
Why This Approach
The Inevitability of the Choice
The authors faced a significant hurdle in synthesizing contrast-enhanced MRI (CEMRI) from non-contrast MRI (NCMRI), particularly for brain tumors. They explicitly state that this task is "highly ill-posed" because NCMRIs provide only ambiguous evidence for enhanced regions. Traditional state-of-the-art (SOTA) methods, including standard CNNs, basic diffusion models like Palette, and even more advanced diffusion models like I2SB, were found to produce "noticeable false positive and false negative enhancement results." This was especially problematic for tumor regions, where these models failed to capture the intricate morphology of tumor subcomponents.
The critical realization that led to the D³M approach was a fundamental reinterpretation of these enhancement errors. Instead of viewing them as large, difficult-to-correct intensity errors, the authors recognized them as an "incorrect interpretation of tumor subcomponents," where enhanced regions were misinterpreted as non-enhanced, and vice versa. This shift in perspective made it clear that the problem could be more effectively addressed through geometric correction via spatial deformation. The authors understood that while intensity errors are typically large and challenging, the necessary geometric corrections (small displacements) are "relatively small and more manageable." This insight made a deformation-driven approach not just an improvement, but the only viable solution to overcome the inherent ill-posedness and the specific failure modes of existing methods in handling complex tumor structures.
Comparative Superiority
The D³M method achieves qualitative superiority through several structural advantages that go beyond mere performance metrics. The core innovation lies in its ability to address enhancement errors geometrically rather than as intensity errors. This is a profound structural advantage because correcting small spatial misalignments is inherently more stable and effective than trying to fix large, erroneous intensity values.
Specifically, the Multi-step Spatial Deformation Module (MSSDM) is a key differentiator. Unlike traditional post-processing deformation, MSSDM is tightly integrated within the stepwise denoising process of the diffusion model. This integration is crucial for two reasons: first, it prevents the accumulation of severe errors that would be difficult to correct later; and second, it promotes the joint optimization of both image generation and geometric correction. This means the model learns to synthesize the image while simultaneously correcting its geometric accuracy at each step, leading to a more robust and precise output.
Furthermore, the Dual-stream Image-Mask Decoder (DSIMD) provides an auxiliary task of segmenting the enhanced tumor. This isn't just an add-on; it's a structural enhancement that improves the model's fundamental "understanding of contrast enhancement." By jointly producing intermediate enhanced images and masks, the DSIMD provides explicit, high-level guidance to the MSSDM for spatial deformation. This dual-stream processing ensures that the geometric corrections are informed by a clear semantic understanding of the tumor's boundaries and enhanced regions, making the overall synthesis qualitatively superior, especially in preserving the intricate morphology of tumors. The ablation studies confirm that both MSSDM and DSIMD contribute significantly to the improved performance, particularly in tumor areas, underscoring their structural importance.
Alignment with Constraints
The D³M approach demonstrates a remarkable "marriage" between the problem's harsh requirements and its unique solution properties. The primary constraint, as identified in the problem definition, is the "highly ill-posed" nature of CEMRI synthesis from NCMRIs, which leads to "false positive and false negative enhancement" and a failure to capture "intricate morphology of tumor subcomponents."
D³M aligns perfectly with these constraints by:
1. Reinterpreting Errors: The solution's core idea of reformulating enhancement errors as geometric misinterpretations directly tackles the ill-posedness. Instead of struggling with large, unmanageable intensity errors, D³M focuses on smaller, more tractable geometric corrections. This property is uniquely suited to the challenge of ambiguous evidence from NCMRIs.
2. Multi-step Geometric Correction (MSSDM): The integration of MSSDM within the diffusion process allows for incremental, step-wise geometric adjustments. This directly addresses the problem of false positive and false negative enhancements by "displacing enhanced regions to remove" these errors. The ability to correct errors continuously throughout the denoising process is a perfect fit for refining the intricate and often subtle details of tumor morphology, which previous methods struggled with.
3. Enhanced Understanding (DSIMD): The auxiliary task of tumor segmentation via DSIMD provides the model with a deeper, semantic understanding of what constitutes an enhanced tumor. This explicit guidance is crucial for accurately synthesizing the "intricate morphology of tumor subcomponents," ensuring that the geometric corrections are biologically plausible and clinically relevant. This dual-stream approach ensures the model doesn't just generate an image, but one that respects the underlying anatomical and pathological structures, a critical requirement for medical imaging.
Rejection of Alternatives
The paper implicitly, yet strongly, rejects alternative approaches by highlighting their fundamental limitations in addressing the specific challenges of CEMRI synthesis from NCMRIs. While the authors do not provide a direct "rejection statement" for each alternative, they establish a clear rationale for why existing methods, including popular ones like GANs and basic diffusion models, are insufficient.
The core reasoning for rejecting alternatives stems from their inability to effectively handle the "highly ill-posed" nature of the problem and the resulting "noticeable false positive and false negative enhancement results," particularly concerning the "intricate morphology of tumor subcomponents." Methods like Pix2Pix (GAN-based) and Palette (a basic diffusion model) are fundamentally designed to learn mappings or generate images based on intensity distributions. Their primary limitation, as implied by the paper, is that they treat enhancement errors predominantly as intensity errors. As the authors explain, these intensity errors are "typically large and challenging to correct."
The paper's comparative analysis (Table 1) further reinforces this rejection. D³M consistently outperforms Pix2Pix, ResViT (a multimodal model combining transformers and CNNs with adversarial learning), Palette, and I2SB (a Schrödinger bridge diffusion model), especially within the critical tumor regions. This empirical evidence, coupled with the theoretical argument that existing methods fail to capture intricate tumor morphology due to their intensity-based error handling, serves as a compelling rejection of these alternatives. The D³M's geometric correction paradigm is presented as the necessary departure from these prior approaches, which were simply not equipped to resolve the specific type of misinterpretation errors inherent in this challenging medical image synthesis task.
Mathematical & Logical Mechanism
The Master Equation
The core of the D³M model, particularly during the inference phase where the contrast-enhanced MRI (CEMRI) is synthesized, is the iterative denoising and geometric correction step. This process is encapsulated by the following equation, which updates the image from a noisy state at time $t$ to a less noisy, geometrically corrected state at time $t-1$:
$$x_{t-1} = \phi_{\hat{u}_t} \left( \sqrt{\bar{\alpha}_{t-1}} \hat{x}_0^{(t)} + \sqrt{1-\bar{\alpha}_{t-1}} \hat{\epsilon}_{i,t} \right)$$
This equation represents a modified deterministic reverse step of a Denoising Diffusion Implicit Model (DDIM), where the predicted original image $\hat{x}_0^{(t)}$ and predicted noise $\hat{\epsilon}_{i,t}$ are first estimated from the current noisy image $x_t$ and the model's velocity predictions, and then spatially deformed by $\phi_{\hat{u}_t}$ before being combined to form the next, less noisy image $x_{t-1}$.
Term-by-Term Autopsy
Let's dissect the master equation and its underlying components:
- $x_{t-1}$: This represents the estimated CEMRI image at the previous time step $t-1$. It is the output of the current reverse diffusion step, having undergone both denoising and geometric correction. Its physical role is to progressively reconstruct the desired CEMRI from noise.
- $\phi_{\hat{u}_t}(\cdot)$: This is the spatial deformation operator. It applies a geometric transformation to its input, warping the image content according to the deformation field $\hat{u}_t$. The author used this operator instead of simple arithmetic operations because the problem is framed as a geometric misinterpretation of tumor subcomponents, requiring spatial adjustment rather than just intensity correction.
- $\hat{u}_t$: This is the predicted deformation field at time step $t$. It is a 2D vector field (for 2D slices) that specifies how each pixel should be displaced. Its physical role is to geometrically correct false positive and false negative enhancements in the synthesized image. It's estimated by the Deformation Estimation Module $U(\cdot)$ within the Multi-Step Spatial Deformation Module (MSSDM), taking the intermediate mask estimate $\hat{m}_0^{(t)}$ as input.
- $\hat{m}_0^{(t)}$: This is an intermediate estimate of the original (ground truth) enhanced tumor mask at time $t$. It's computed from the noisy mask $m_t$ and the predicted mask velocity $\hat{v}_{m,t}$ using the formula:
$$\hat{m}_0^{(t)} = \sqrt{\bar{\alpha}_t} m_t - \sqrt{1-\bar{\alpha}_t} \cdot \hat{v}_{m,t}$$
Its logical role is to provide a clean, uncorrupted representation of the tumor's enhanced regions, which is crucial for guiding the deformation field estimation. The subtraction here is part of reversing the forward diffusion process, effectively "denoising" the mask.
- $\sqrt{\bar{\alpha}_{t-1}}$: This is a scaling factor for the predicted original image component. It's derived from the noise schedule parameters and dictates the weight of the "signal" (predicted original image) in the reconstruction. The square root is standard in diffusion models to handle variances.
- $\hat{x}_0^{(t)}$: This is the model's prediction of the original (non-noisy) CEMRI image, estimated from the current noisy image $x_t$ and the predicted image velocity $\hat{v}_{i,t}$ at time step $t$. Its physical role is to represent the underlying clean image that the diffusion process is trying to recover. It is derived as:
$$\hat{x}_0^{(t)} = \frac{\sqrt{\alpha_t} x_t - \sqrt{1-\bar{\alpha}_t} \hat{v}_{i,t}}{\sqrt{\alpha_t \bar{\alpha}_t} + 1-\bar{\alpha}_t}$$
The combination of addition/subtraction and scaling factors is a direct consequence of inverting the forward diffusion process and relating the predicted velocity to the predicted original image. - $\sqrt{1-\bar{\alpha}_{t-1}}$: This is another scaling factor, applied to the predicted noise component. It represents the weight of the "noise" in the reconstruction. Like $\sqrt{\bar{\alpha}_{t-1}}$, its form is derived from the variance schedule.
- $\hat{\epsilon}_{i,t}$: This is the model's prediction of the Gaussian noise component that was added to the original image to produce $x_t$. Its physical role is to represent the random fluctuations that need to be removed. It is derived from $x_t$ and the predicted image velocity $\hat{v}_{i,t}$ as:
$$\hat{\epsilon}_{i,t} = \frac{\sqrt{1-\bar{\alpha}_t} x_t + \sqrt{\bar{\alpha}_t} \hat{v}_{i,t}}{\sqrt{\alpha_t \bar{\alpha}_t} + 1-\bar{\alpha}_t}$$
Again, the arithmetic operations are part of the mathematical inversion of the diffusion process. - $x_t$: This is the noisy CEMRI image at the current time step $t$, serving as the input to the current denoising step.
- $\hat{v}_{i,t}$: This is the predicted image velocity term at time step $t$. It's the primary output of the Dual-Stream Image-Mask Decoder (DSIMD) $D(\cdot)$, which takes the noisy image $x_t$, noisy mask $m_t$, conditional NCMRIs $c$, and time step $t$ as input:
$$(\hat{v}_{i,t}, \hat{v}_{m,t}) = D(E(x_t, m_t, c, t), t)$$
Its logical role is to provide a more stable and efficient target for the model to learn, as opposed to directly predicting noise or the denoised image. - $m_t$: This is the noisy enhanced tumor mask at the current time step $t$. It's processed alongside $x_t$ to guide the mask velocity prediction.
- $c$: These are the conditional images, specifically the non-contrast magnetic resonance images (NCMRIs). They provide crucial anatomical context to the model, guiding the synthesis of the CEMRI. They are concatenated with the noisy inputs before being fed into the encoder.
- $t$: This represents the current time step in the diffusion process, ranging from $T$ (pure noise) down to $0$ (clean image). It's fed into the network to inform the model about the current noise level.
- $E(\cdot)$: This is the encoder network, based on PixelCNN++ with a Wide ResNet backbone. Its role is to extract relevant features from the noisy image, mask, and conditional NCMRIs.
- $D(\cdot)$: This is the Dual-Stream Image-Mask Decoder (DSIMD), which takes the encoded features and the time step $t$ to jointly produce the predicted image velocity $\hat{v}_{i,t}$ and mask velocity $\hat{v}_{m,t}$. The dual-stream design allows for separate handling of image and mask information, improving the model's understanding of contrast enhancement.
- $\bar{\alpha}_t = \prod_{s=1}^t \alpha_s$: This is the cumulative product of the noise schedule parameters $\alpha_s$ up to time $t$. It determines the overall scaling of the original image component in the forward diffusion process. The product is used because variances add up in independent Gaussian noise additions.
- $\alpha_t$: This is a parameter from the noise schedule at time step $t$, dictating the amount of noise added or removed at each step.
Step-by-Step Flow
Imagine a single abstract data point, representing a voxel in the brain, as it undergoes transformation during the inference (synthesis) process. The goal is to generate a clean, contrast-enhanced MRI ($x_0$) from an initial state of pure noise ($x_T$).
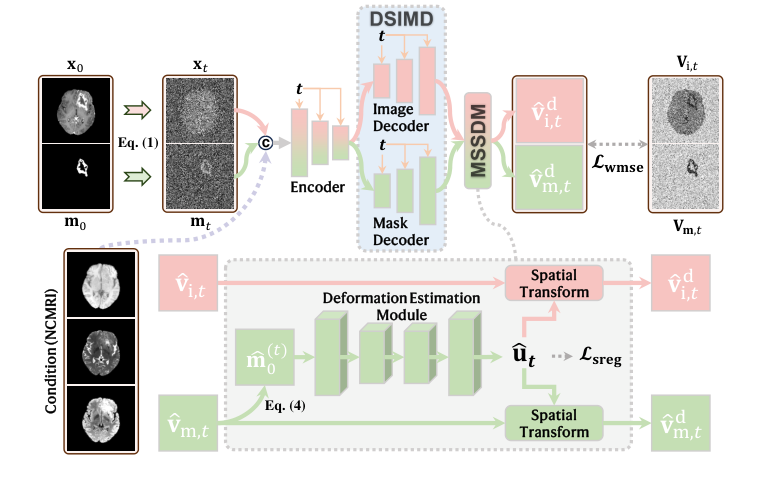 Figure 1. An overview of the network architecture of D3M
Figure 1. An overview of the network architecture of D3M
- Initialization: The process begins at the largest time step, $t=T$. We start with a completely noisy image $x_T$ and a noisy mask $m_T$, both essentially random Gaussian noise.
- Feature Extraction (Encoder): At each step $t$ (starting from $T$ and going down to $1$), the current noisy image $x_t$, the noisy mask $m_t$, the conditional non-contrast MRIs $c$, and the current time step $t$ are fed into the Encoder $E(\cdot)$. This encoder acts like a sophisticated filter, extracting meaningful features that represent the underlying structure and enhancement patterns, even amidst the noise.
- Velocity Prediction (DSIMD): The features extracted by the encoder are then passed to the Dual-Stream Image-Mask Decoder $D(\cdot)$. This decoder, designed with two parallel streams, processes these features to predict two crucial "velocity" terms: $\hat{v}_{i,t}$ for the image and $\hat{v}_{m,t}$ for the mask. These velocities indicate the direction and magnitude of change needed to move from the noisy state towards the clean original image and mask.
- Intermediate Image and Mask Estimation:
- Using the predicted image velocity $\hat{v}_{i,t}$ and the current noisy image $x_t$, the model mathematically inverts the forward diffusion process to estimate what the original, clean CEMRI image ($\hat{x}_0^{(t)}$) would look like, and what the noise component ($\hat{\epsilon}_{i,t}$) was. This is done using the derived formulas for $\hat{x}_0^{(t)}$ and $\hat{\epsilon}_{i,t}$.
- Similarly, using the predicted mask velocity $\hat{v}_{m,t}$ and the current noisy mask $m_t$, an intermediate estimate of the original, clean enhanced tumor mask ($\hat{m}_0^{(t)}$) is computed (as per Eq 4). This mask is critical for understanding tumor subcomponents.
- Deformation Field Estimation (MSSDM): The intermediate mask estimate $\hat{m}_0^{(t)}$ is then fed into the Deformation Estimation Module $U(\cdot)$, which is part of the Multi-Step Spatial Deformation Module (MSSDM). This module analyzes the estimated mask to identify regions of false positive or false negative enhancement and calculates a deformation field $\hat{u}_t$. This field specifies how pixels need to be shifted to geometrically correct these enhancement errors.
- Geometric Correction (Spatial Deformation): The estimated original image $\hat{x}_0^{(t)}$ and noise estimate $\hat{\epsilon}_{i,t}$ are then passed through the spatial deformation operator $\phi_{\hat{u}_t}$. This operator warps these images according to the deformation field $\hat{u}_t$, effectively "moving" enhanced regions to their correct locations or removing spurious ones. This is a crucial step that distinguishes D³M, as it corrects errors geometrically rather than just by intensity.
- Reverse Diffusion Step: Finally, the deformed $\hat{x}_0^{(t)}$ and $\hat{\epsilon}_{i,t}$ are combined using the DDIM reverse step (the master equation). This step effectively removes a small amount of noise from the image, producing a new, less noisy, and geometrically corrected image $x_{t-1}$.
- Iteration: This entire process repeats, with $x_{t-1}$ becoming the new $x_t$ for the next step, until $t$ reaches $0$. At $t=0$, the final synthesized CEMRI image $x_0$ is obtained, representing the model's best estimate of the contrast-enhanced image with accurate tumor morphology.
This sequential, iterative process allows the model to gradually refine the image, denoising it while simultaneously correcting geometric distortions related to tumor enhancement.
Optimization Dynamics
The D³M mechanism learns and converges by iteratively adjusting its internal parameters (weights of the neural networks) based on a carefully constructed loss function. This process is driven by the interplay of two main components:
-
Loss Function: The model is trained to minimize a combined loss, which consists of two parts:
- Weighted Mean Squared Error Loss ($\mathcal{L}_{wmse}$): This is a standard loss in diffusion models, typically measuring the discrepancy between the model's predictions (e.g., predicted noise $\hat{\epsilon}_{i,t}$ or predicted velocity $\hat{v}_{i,t}$) and the true values. The paper states it's applied to the "synthesis result at each step," implying it guides the model to produce accurate CEMRI images. The "weighted" aspect means that different parts of the image or different time steps might contribute differently to the overall loss, potentially focusing more on critical regions like tumors. This loss shapes the landscape by creating "valleys" where the model's predictions closely match the ground truth, encouraging accurate image synthesis.
- Deformation Smoothness Regularization ($\mathcal{L}_{sreg}$): This term is applied to the deformation field $\hat{u}_t$ at each step. It typically penalizes large or abrupt changes in the deformation field, often by computing an L2 norm of its spatial gradients. Its purpose is to ensure that the geometric corrections are smooth and physically plausible, preventing the model from generating unrealistic or jagged deformations. This regularization term acts as a "penalty" that flattens out overly complex or noisy regions in the loss landscape related to deformation, guiding the model towards more stable and interpretable transformations. The author used this to prevent the model from overfitting to noise or creating artifacts in the deformation.
-
Gradient-Based Optimization: During training, the combined loss is calculated, and then gradients of this loss with respect to all trainable parameters in the Encoder $E(\cdot)$, Dual-Stream Image-Mask Decoder $D(\cdot)$, and Deformation Estimation Module $U(\cdot)$ are computed using backpropagation. These gradients indicate the direction and magnitude by which each parameter should be adjusted to reduce the loss.
-
Optimizer: The Adam optimizer [12] is employed to update the model's parameters. Adam is an adaptive learning rate optimization algorithm that efficiently adjusts the learning rate for each parameter based on estimates of first and second moments of the gradients. This helps the model navigate the complex loss landscape more effectively and converge faster. The learning rate is set to $8 \times 10^{-5}$, and a batch size of 16 is used, meaning parameters are updated after processing 16 image-mask pairs.
-
Iterative State Updates and Convergence: The model undergoes 200,000 training iterations. In each iteration, a batch of data is processed, the loss is computed, and the parameters are updated. This iterative process allows the model to gradually learn the intricate relationships between noisy inputs, conditional information, and the desired clean, geometrically corrected CEMRI and mask. The loss landscape, shaped by both $\mathcal{L}_{wmse}$ and $\mathcal{L}_{sreg}$, guides the model towards a minimum where it can accurately predict velocities and deformation fields. Convergence is achieved when the model's performance on validation data no longer significantly improves, indicating that it has learned a stable mapping for synthesizing high-quality CEMRIs with precise geometric correction, as evidenced by the superior PSNR and SSIM scores. The joint optimization of image generation and geometric correction, facilitated by the tight integration of MSSDM within the denoising process, helps prevent error accumulation and promotes a more robust convergence.
Results, Limitations & Conclusion
Experimental Design & Baselines
To rigorously validate their deformation-driven diffusion model (D³M), the authors conducted extensive experiments on two publicly available datasets: BraSyn [13] and BraTS-PEDs [11]. These datasets are rich, comprising brain magnetic resonance images from 1,470 and 307 patients, respectively, all diagnosed with brain tumors. Crucially, they include aligned T1-weighted, T2-weighted, FLAIR, and the target contrast-enhanced T1-weighted (CEMRI) images. For training, the model leveraged auxiliary information in the form of manually annotated and clinically reviewed enhanced tumor masks, available for 1,251 patients in BraSyn and 216 in BraTS-PEDs. The datasets were split into training, validation, and test sets to ensure unbiased evaluation, with 1,001/250/219 patients for BraSyn and 173/43/91 for BraTS-PEDs, respectively.
The "victims" (baseline models) against which D³M was ruthlessly tested included a diverse range of state-of-the-art image synthesis methods. These encompassed:
- Pix2Pix [10]: A foundational GAN-based image-to-image translation model utilizing a CNN architecture.
- ResViT [4]: A more recent multimodal medical image synthesis model that ingeniously combines vision transformers with convolutional operators and adversarial learning.
- Palette [18]: A representative basic diffusion model for image synthesis.
- I2SB [14]: A Schrödinger bridge diffusion model, which is an advanced variant that improves upon conventional diffusion models.
To ensure a fair fight, all competing methods were trained, validated, and tested using the exact same data splits as D³M. Furthermore, the authors went a step further to level the playing field by also providing the enhanced tumor masks as an auxiliary task during training for the baseline methods, mirroring D³M's approach. This meticulous experimental design aimed to isolate the impact of D³M's core architectural innovations.
The performance was quantitatively assessed using two widely accepted metrics in image quality assessment: Peak Signal-to-Noise Ratio (PSNR) and Structural Similarity Index Measure (SSIM), calculated between the synthesized and real CEMRIs. Recognizing the critical importance of tumor regions, these metrics were also specifically computed within these areas. To delineate tumor regions for test images, an nnU-Net segmentation model [9] was trained on the tumor annotations from the training data. Statistical significance of the improvements was determined using the Wilcoxon signed-rank test, with a stringent threshold of $p < 0.001$. Images were normalized by clipping intensity values between the 0.5th and 99.5th percentiles and rescaling to [0,1], with an input size of 256 × 256, trained for 200,000 iterations using the Adam optimizer [12] with a batch size of 16 and a learning rate of $8 \times 10^{-5}$.
What the Evidence Proves
The evidence unequivocally proves the efficacy of D³M's novel approach to CEMRI synthesis, particularly its core mechanism of geometrically correcting enhancement errors through spatial deformation. The authors' hypothesis that reformulating enhancement errors as incorrect interpretations of tumor subcomponents, rather than mere intensity errors, allows for more manageable geometric correction, is strongly supported by both qualitative and quantitative results.
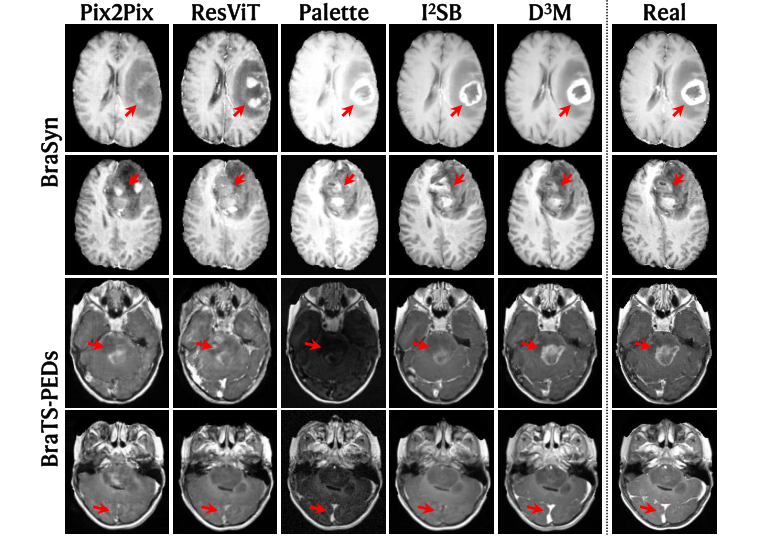 Figure 2. Examples of synthesis results, shown together with the real CEMRI for refer- ence. Note the tumor and vessel regions highlighted by arrows for comparison
Figure 2. Examples of synthesis results, shown together with the real CEMRI for refer- ence. Note the tumor and vessel regions highlighted by arrows for comparison
-
Visual Superiority (Fig. 2): The qualitative results presented in Figure 2 offer compelling visual proof. D³M's synthesized CEMRIs are strikingly "more consistent with the real image" compared to all baseline methods. Crucially, within the intricate tumor and vessel regions (highlighted by red arrows), D³M accurately reproduces the enhancement patterns that closely match the ground truth. In stark contrast, competing methods frequently exhibit "noticeable false positive and/or false negative enhancement," generating either spurious enhanced regions or failing to enhance areas that should be. This visual fidelity, especially in complex pathological structures, is a direct testament to D³M's ability to capture and correct subtle geometric discrepancies.
-
Quantitative Dominance (Table 1): The quantitative metrics provide hard, statistical evidence. D³M consistently achieved the highest PSNR and SSIM values across both the BraSyn and BraTS-PEDs datasets, not only for the "Whole Image" but, more importantly, for the "Tumor Region." For instance, on BraSyn, D³M scored a PSNR of $25.11 \pm 3.33$ and SSIM of $90.95 \pm 3.86$ for the whole image, and a PSNR of $17.33 \pm 4.56$ and SSIM of $73.21 \pm 16.22$ for the tumor region. These figures represent a clear improvement over all baselines. The statistical analysis, using the Wilcoxon signed-rank test, confirmed that these improvements are "highly statistically significant ($p < 0.001$)," leaving no room for doubt about D³M's superior performance. This quantitative edge, particularly within the diagnostically critical tumor regions, underscores that D³M's mechanism effectively mitigates the "ill-posed" nature of CEMRI synthesis.
-
Ablation Study Validation (Table 2): The ablation studies on the BraSyn dataset provide direct, component-level proof of D³M's architectural innovations.
- Benefit of MSSDM: When the Multi-Step Spatial Deformation Module (MSSDM) was removed, leading to direct use of DSIMD's output for velocity terms, both PSNR and SSIM decreased significantly, "especially in the tumor area." This directly validates that MSSDM's geometric correction, which deforms the synthesized image to adjust enhancement, is a critical component for accurate tumor representation. It confirms that correcting enhancement through spatial deformation, rather than just intensity adjustments, is indeed beneficial.
- Benefit of DSIMD: Further removing the Dual-Stream Image-Mask Decoder (DSIMD) and replacing it with a single decoder resulted in an even more pronounced drop in PSNR and SSIM. This demonstrates the indispensable role of DSIMD in jointly producing intermediate enhanced images and masks. This dual-stream approach evidently provides the necessary guidance for MSSDM, enhancing the model's understanding of contrast enhancement and enabling more precise geometric corrections.
In sum, the experimental architecture, the defeat of strong baseline models, and the meticulous ablation studies provide definitive and undeniable evidence that D³M's core mechanism—geometric correction of enhancement errors via spatial deformation guided by a dual-stream image-mask decoder—is a highly effective and statistically significant advancement in synthesizing high-quality CEMRIs, particularly for brain tumors.
Limitations & Future Directions
While D³M presents a significant leap forward in CEMRI synthesis, particularly for brain tumors, it's important to acknowledge that no scientific endeavor is without its boundaries and avenues for future exploration. The paper itself, while not explicitly listing a "Limitations" section, implicitly points to the inherent complexity of the problem, noting that CEMRI synthesis from NCMRIs is "highly ill-posed" and that existing methods struggle with "intricate morphology of tumor subcomponents." While D³M addresses these challenges effectively, the absolute PSNR and SSIM values, even for the best-performing model, suggest there's still a gap between synthetic and real images, indicating room for further refinement. The geometric correction, while powerful, is described as "relatively small," implying it's a targeted refinement rather than a complete overhaul of the synthesis process.
Looking ahead, D³M's findings open up several exciting and critical discussion topics for future development:
-
Generalizability Across Pathologies and Anatomies: The current success is demonstrated on brain tumors. A crucial next step is to investigate D³M's performance and adaptability to other pathologies (e.g., inflammation, metastases) and different anatomical regions (e.g., liver, prostate, breast). Each organ and disease presents unique challenges in terms of contrast enhancement patterns and anatomical variability. What architectural modifications or training strategies would be necessary to maintain or even improve performance in these diverse contexts?
-
Clinical Integration and Regulatory Pathways: The ultimate goal of such research is clinical utility. How can synthetic CEMRIs be rigorously validated for diagnostic accuracy and treatment planning in real-world clinical settings? What level of agreement with actual CEMRIs is deemed acceptable by clinicians and regulatory bodies? This involves not just image quality metrics but also clinical outcome studies. Furthermore, the ethical implications of using AI-generated images for patient care, including issues of liability and patient trust, need careful consideration and robust regulatory frameworks.
-
Uncertainty Quantification and Explainability: In medical imaging, confidence in a diagnosis is paramount. While D³M generates high-quality images, quantifying the uncertainty associated with the synthesized enhancement, especially in critical tumor margins, would be invaluable. Techniques for uncertainty estimation could provide clinicians with a measure of reliability for the synthetic images. Additionally, improving the explainability of the deformation fields—understanding why the model applies specific geometric corrections—could foster greater trust and interpretability for medical professionals.
-
Computational Efficiency and Real-time Applications: Diffusion models, while powerful, can be computationally intensive, particularly for 3D volumes and multi-step processes. For clinical adoption, especially in time-sensitive scenarios or for large-scale data processing, optimizing D³M for faster inference is essential. Exploring advanced sampling techniques, model distillation, or hardware acceleration could significantly reduce synthesis time without compromising quality. This would enable more practical integration into clinical workflows.
-
Longitudinal Monitoring and Disease Progression: CEMRIs are frequently used for longitudinal monitoring of tumor response to treatment. Can D³M consistently synthesize CEMRIs that accurately reflect subtle changes in tumor size, morphology, and enhancement over time? Maintaining consistency and sensitivity to progression or regression across multiple time points is a complex challenge that could greatly benefit from D³M's geometric correction capabilities.
-
Multi-modal and Multi-sequence Fusion: The current NCMRIs include T1, T2, and FLAIR. Future work could explore incorporating additional non-contrast sequences, such as diffusion-weighted imaging (DWI) or perfusion imaging, which provide complementary physiological information. Could a richer input space further enhance the model's understanding of tissue properties and lead to even more accurate and robust CEMRI synthesis?
These discussion points highlight that while D³M has made significant strides, the journey towards fully realizing the potential of synthetic medical imaging is ongoing, requiring interdisciplinary collaboration and continuous innovation.
