Prompt-DAS: Annotation-Efficient Prompt Learning for Domain Adaptive Semantic Segmentation of Electron Microscopy Images
Prompt DAS adapts AI to segment tiny cell parts in electron microscope images, offering flexible, efficient, and interactive annotation.
Background & Academic Lineage
The field of semantic segmentation, which involves precisely outlining and categorizing every pixel in an image, has seen remarkable advancements thanks to deep neural networks like Convolutional Neural Networks (CNNs) and Vision Transformers (ViTs). This progress has been particularly impactful in analyzing Electron Microscopy (EM) images, which are crucial for studying subcellular organelles such as mitochondria in cancer research and biology.
However, this powerful technology faced two significant hurdles. First, training these sophisticated models demands an enormous amount of pixel-wise annotations. Imagine having to meticulously draw the exact boundary around every single mitochondrion in thousands of ultra-detailed EM images – it's an incredibly time-consuming, expensive, and labor-intensive task that often requires highly specialized experts. This high annotation burden made it impractical for large-scale applications.
Second, models trained on one set of EM images (the "source domain") often perform poorly when applied to new EM images (the "target domain") that come from different microscopes or tissue types. This phenomenon, known as "domain shift," means a model trained on rat brain images might struggle with human liver images, even if both contain mitochondria.
To address the annotation burden and domain shift, researchers turned to "Domain Adaptation" (DA). Early attempts included Unsupervised Domain Adaptation (UDA), which assumes no annotations on the target domain. While appealing, UDA methods often yielded relatively low performance on complex segmentation tasks, limiting their real-world utility. Weakly Supervised Domain Adaptation (WDA) emerged as a more practical alternative, leveraging "sparse points" (just a few dots indicating an object's location) as cheap, weak labels on the target domain to boost performance with minimal annotation effort. While better, WDA still required some manual input and wasn't always flexible enough for varying annotation scenarios.
More recently, the advent of "prompt-driven" foundation models like the Segment Anything Model (SAM) revolutionized segmentation for natural images. SAM, pre-trained on billions of images, could segment objects based on simple "prompts" like a single click (a point), a bounding box, or a rough scribble. This opened the door for interactive segmentation, where users could guide the model.
However, SAM itself had critical limitations when applied to the medical and EM imaging domain. It struggled with domain shifts in medical images and often showed low performance, especially with point prompts, due to its lack of specific medical knowledge, ambiguous biological boundaries, and complex shapes of organelles. Crucially, SAM requires a prompt for each individual object instance. For EM images teeming with hundreds or thousands of tiny organelles, providing a prompt for every single one is just as impractical as pixel-wise annotation. Furthermore, SAM's performance with point prompts, specifically, was often suboptimal in medical contexts.
These limitations—the prohibitive cost of pixel-wise annotations, the poor generalization across different EM image datasets, and the shortcomings of existing DA methods and even powerful foundation models like SAM in the medical domain—collectively forced the authors to develop Prompt-DAS. Their motivation was to create a flexible, annotation-efficient, promptable transformer model specifically for domain-adaptive semantic segmentation of EM images that could overcome these challenges by effectively utilizing sparse point prompts during both training and testing, and performing well across UDA, WDA, and interactive segmentation scenarios.
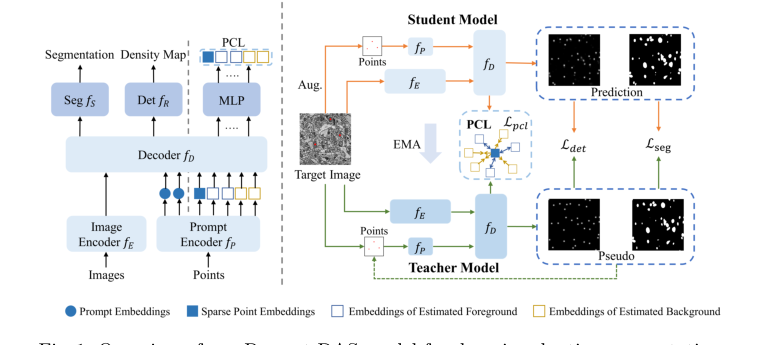 Figure 1. Overview of our Prompt-DAS model for domain adaptive segmentation
Figure 1. Overview of our Prompt-DAS model for domain adaptive segmentation
Domain Term Analogies
- Semantic Segmentation: Imagine you have a coloring book with a picture of a garden. Semantic segmentation is like coloring all the "flowers" red, all the "leaves" green, and all the "soil" brown. You're not just drawing a box around each flower; you're coloring every single pixel that belongs to a flower, a leaf, or the soil, based on its category.
- Domain Adaptation (DA): Think of a student who learns to identify different types of cars in sunny California (the "source domain"). Now, this student moves to snowy Alaska (the "target domain") and needs to identify cars there. Instead of forgetting everything and starting over, domain adaptation is like the student using their existing knowledge of cars but adjusting it to account for the snow, ice, and different lighting conditions. They adapt their learning to the new environment.
- Electron Microscopy (EM) Images: Picture looking at the tiny, intricate gears inside a watch. A regular camera uses light to take a picture. An electron microscope, however, uses a beam of electrons instead of light to "see" things that are incredibly small, like the internal structures of a cell, at a much higher magnification and detail than a regular microscope. So, EM images are like super-detailed, high-contrast black-and-white photographs of the microscopic world.
- Promptable Learning: Imagine you have a very talented artist who can draw anything. Instead of just saying "draw a house," which is vague, you give them a "prompt"—like pointing to a specific spot on the paper and saying "draw a house here," or drawing a rough outline and saying "fill this shape with a house." The "prompt" is a small, specific hint (like a point or a box) that guides the artist (the AI model) to perform a complex task (drawing/segmenting) exactly where and how you want it.
- Pseudo-labeling: Consider a teacher giving you a quiz. Some questions have answers provided (labeled data), but most don't. You try to answer the questions without answers, and for the ones you're very confident about, you mark them as correct. Then, you use these "self-graded" answers (pseudo-labels) to study even more, as if they were real answers. In AI, a "teacher model" generates these confident "self-graded" answers for a "student model" to learn from, helping it learn from unlabeled data.
Notation Table
| Notation | Type | Description |
|---|---|---|
| $D_s$ | Variable | Source domain dataset, consisting of images and their full pixel-wise labels. |
| $D_t$ | Variable | Target domain dataset, consisting of images and sparse point labels. |
| $x^s, x^t$ | Variable | Input images from the source and target domains, respectively. |
| $y^s$ | Variable | Full pixel-wise ground truth labels for source images. |
| $c^t$ | Variable | Ground truth point labels for a few object instances in target images. |
| $\hat{c}^t$ | Variable | Binary dot label map, where 1 indicates an annotated sparse point. |
| $d$ | Variable | Density map, derived from point labels by convolution with a Gaussian kernel. |
| $k_\sigma$ | Parameter | Gaussian kernel used to generate the density map. |
| $f_e$ | Model Component | Image encoder, extracts features from input images. |
| $f_p$ | Model Component | Point prompt encoder, processes input point prompts. |
| $f_D$ | Model Component | Multitask decoder, integrates image and prompt features. |
| $f_s$ | Model Component | Semantic segmentation head, outputs segmentation predictions. |
| $f_r$ | Model Component | Regression-based center-point detection head, outputs point detections. |
| $M$ | Parameter | Number of points provided as input to the prompt encoder. |
| $L_{det}$ | Variable | Detection loss, measures accuracy of center-point detection. |
| $L_{seg}$ | Variable | Segmentation loss, measures accuracy of semantic segmentation. |
| $L_{pcl}$ | Variable | Prompt-guided contrastive loss, enhances feature discriminability. |
| $F_R$ | Model Component | Full detection network, composed as $f_r \circ f_D \circ f_e$. |
| $F_S$ | Model Component | Full segmentation network, composed as $f_s \circ f_D \circ f_e$. |
| $MSE$ | Variable | Mean Square Error loss function. |
| $CE$ | Variable | Cross-Entropy loss function. |
| $\hat{d}^t$ | Variable | Predicted density map for the target domain, used for pseudo-labeling. |
| $\hat{y}^t$ | Variable | Pseudo labels generated by the teacher model for target segmentation. |
| $n_s$ | Parameter | Number of randomly sampled center points used as training prompts for source data. |
| $z^t$ | Variable | Feature embedding derived from a target domain point $p^t$. |
| $\phi$ | Model Component | MLP (Multi-Layer Perceptron) layer used before contrastive learning. |
| $N_q$ | Variable | Number of foreground prompt embeddings. |
| $N_n$ | Variable | Number of background prompt embeddings. |
| $\mu^s$ | Variable | Average embedding of sparse point prompts. |
| $\tau$ | Parameter | Temperature parameter for the contrastive loss, controlling the sharpness of feature separation. |
| $\delta_f$ | Parameter | Confidence threshold for selecting foreground points for pseudo-labeling. |
| $\delta_b$ | Parameter | Confidence threshold for selecting background points for pseudo-labeling. |
Problem Definition & Constraints
Core Problem Formulation & The Dilemma
The Starting Point (Input/Current State):
The authors begin with a source domain $\mathcal{D}^s = \{(x^s, y^s)\}$ containing electron microscopy (EM) images with full, pixel-wise ground-truth labels. They also have a target domain $\mathcal{D}^t = \{(x^t, \bar{c}^t)\}$, which consists of images from a different distribution (e.g., different tissue types or microscopy techniques) where only sparse, point-based labels $\bar{c}^t$ are available for a small subset of organelle instances.
The Desired Endpoint (Output/Goal State):
The goal is to develop a robust, "promptable" segmentation framework that can accurately segment all organelle instances in the target domain. The model must be flexible enough to function under three distinct scenarios:
1. Unsupervised Domain Adaptation (UDA): Where $M=0$ points are provided on the target training data.
2. Weakly Supervised Domain Adaptation (WDA): Where $M > 0$ sparse points are provided as training prompts.
3. Interactive Segmentation: Where the model can accept user-provided point prompts during the testing phase to refine or correct segmentation results.
The Dilemma & Constraints:
The primary dilemma is the "annotation-performance trade-off." While deep neural networks (like U-Net or Vision Transformers) achieve high accuracy, they are notoriously data-hungry and require expensive, expert-level pixel-wise annotations. When these models are trained on one domain and applied to another, they suffer from significant performance degradation due to domain shifts.
The authors hit several "harsh walls" that make this problem difficult:
* Domain Shift: EM images vary wildly in appearance based on the specific microscopy technique used, rendering standard pre-trained models (like the original SAM) ineffective without significant adaptation.
* Label Scarcity: Obtaining pixel-level masks for every organelle in a new EM dataset is labor-intensive and often infeasible for large-scale studies.
* Instance Complexity: Unlike natural images, EM images contain numerous, densely packed, and often ambiguous organelle instances. Standard foundation models like SAM struggle with these because they lack domain-specific medical knowledge and often require a prompt for every individual instance, which is an impractial burden for large-scale biological datasets.
* Computational Constraints: The authors must balance the need for high-resolution feature extraction with the memory limits of modern GPUs (e.g., 24 GB VRAM), while ensuring the model remains efficient enough to be trained from scratch rather than relying on massive, billion-scale pre-training.
To bridge this gap, the authors introduce a multitask framework that treats segmentation as a promptable task, using an auxiliary center-point detection head to generate pseudo-labels and guide the segmentation process. This effectively turns the "sparse point" limitation into a structural advantage, allowing the model to learn discriminative features even when full ground-truth masks are missing.
Why This Approach
The authors of this paper identified a critical bottleneck in electron microscopy (EM) image analysis: while foundation models like SAM (Segment Anything Model) have revolutionized general computer vision, they fail significantly when applied to EM images. This failure stems from a lack of domain-specific knowledge, the presence of complex, ambiguous boundaries in subcellular organelles, and the inability of SAM to segment numerous instances simultaneously without individual prompts.
Why this approach was the only viable solution
The authors realized that traditional "SOTA" methods—including standard UDA (Unsupervised Domain Adaptation) and even fine-tuned foundation models—were insufficient because they either required massive, expert-level pixel-wise annotations or struggled with the "domain shift" inherent in EM imaging. The "aha!" moment occurred when they recognized that the segmentation task could be simplified by treating it as a multitask problem. By coupling a dense segmentation task with a simpler, regression-based center-point detection task, they could generate their own pseudo-prompts. This effectively bypassed the need for expensive manual labeling while maintaining high precision.
Comparative Superiority (The Benchmarking Logic)
The superiority of Prompt-DAS is not merely in its accuracy metrics, but in its structural efficiency:
- Annotation Efficiency: Unlike models that require full pixel-wise masks, Prompt-DAS achieves state-of-the-art results using only 15% sparse point annotations. This reduces the expert labor required by a massive margin.
- Flexibility: While SAM requires a prompt for every single object instance, Prompt-DAS is designed to handle any number of prompts—from zero (UDA) to sparse points (WDA)—and can even perform interactive segmentation during testing.
- Discriminative Learning: The introduction of Prompt-guided Contrastive Learning (PCL) provides a structural advantage. By pulling foreground embeddings closer to ground-truth points and pushing background embeddings away, the model learns a more robust feature representation than standard cross-entropy loss alone could provide.
The "Marriage" of Constraints and Solution
The paper addresses the "harsh requirements" of EM imaging—specifically the scarcity of labels and the high variability of organelle shapes—through a teacher-student framework. By using a teacher model to generate pseudo-labels and employing Non-Maxima Suppression to identify local maxima, the model creates a self-improving loop. This perfectly aligns with the constraint of limited data: the model uses the "easier" detection task to supervise the "harder" segmentation task.
Mathematical & Logical Mechanism
The Mathematical Engine
The core of Prompt-DAS is a multitask learning framework that simultaneously performs center-point detection and semantic segmentation. The "Master Equation" for the system is the combined loss function, which balances these two tasks across source and target domains:
$$\mathcal{L}_{total} = \mathcal{L}_{det} + \mathcal{L}_{seg} + \lambda \mathcal{L}_{pcl}$$
Where $\mathcal{L}_{det}$ and $\mathcal{L}_{seg}$ are defined as:
$$\mathcal{L}_{det} = \frac{1}{|D^s|} \sum_{x^s} MSE(F_R(x^s), d^s) + \frac{1}{|D^t|} \sum_{x^t} MSE(F_R(x^t), \hat{d}^t)$$
$$\mathcal{L}_{seg} = \frac{1}{|D^s|} \sum_{x^s} CE(F_S(x^s), y^s) + \frac{1}{|D^t|} \sum_{x^t} CE(F_S(x^t), \hat{y}^t)$$
Tearing Down the Equations
- $MSE(F_R(x), d)$: This is the Mean Square Error. It measures the pixel-wise difference between the predicted density map and the ground truth (or pseudo-label) density map. It acts as a regression penalty, forcing the model to place "peaks" of high intensity exactly where the center of an organelle is located.
- $CE(F_S(x), y)$: This is the Cross-Entropy loss. It is the standard classification loss for segmentation, penalizing the model when the predicted class probability for a pixel deviates from the ground truth label.
- $F_R$ and $F_S$: These represent the composition of the image encoder $f_E$, the decoder $f_D$, and the respective heads ($f_R$ for detection, $f_S$ for segmentation). The author uses function composition ($\circ$) to denote the flow of data through the shared backbone into specialized task-specific branches.
- $\hat{d}^t$ and $\hat{y}^t$: These are pseudo-labels generated by the teacher model. They are crucial because they provide supervision on the target domain where ground truth is scarce.
Step-by-Step Flow
Imagine a single EM image entering the assembly line:
1. Feature Extraction: The image $x$ passes through the encoder $f_E$, which transforms raw pixels into a high-dimensional feature map.
2. Prompt Injection: If point prompts are available, the prompt encoder $f_P$ converts these coordinates into embeddings, which are then injected into the decoder $f_D$ via cross-attention.
3. Multitask Branching: The decoder $f_D$ splits the flow. One branch ($f_R$) predicts a density map (where the organelles are), and the other ($f_S$) predicts the final segmentation mask.
4. Pseudo-Labeling: The teacher model (an EMA version of the student) looks at the output. If the teacher is confident, it generates a "pseudo-label" that acts as a teacher for the student, effectively guiding it through the unlabelled target domain.
5. Contrastive Refinement: The PCL module takes foreground and background embeddings and uses the contrastive loss to "pull" similar features together and "push" background noise away, ensuring the model doesn't get confused by the complex textures of EM images.
Optimization Dynamics
The model learns through a Mean-Teacher framework. The student model updates its weights via backpropagation using the loss functions above. Simultaneously, the teacher model is updated using an Exponential Moving Average (EMA) of the student's weights. This creates a "stable" teacher that provides consistent, high-quality pseudo-labels, preventing the student from chasing noisy, unstable predictions early in training.
Results, Limitations & Conclusion
Experimental Evidence
The authors ruthlessly tested their model against a variety of "victims," including standard UDA methods (like DAMT-Net) and SAM-based approaches (like WeSAM).
* The Evidence: In Table 1, the results are clear. While SAM and its medical variants (SAM-Med2D) show severely degraded performance on EM images, Prompt-DAS consistently hits higher Dice scores.
* The Proof: The ablation study (Table 2) is the most compelling evidence. It shows that adding detection pseudo-labeling, then segmentation pseudo-labeling, then training prompts, and finally PCL, provides a cumulative, measurable boost in performance. Each component was not just "added"; it was mathematically justified to solve a specific failure mode of the previous step. The model achieves near-supervised performance with only 15% of the annotation effort, which is a massive win for efficiency.
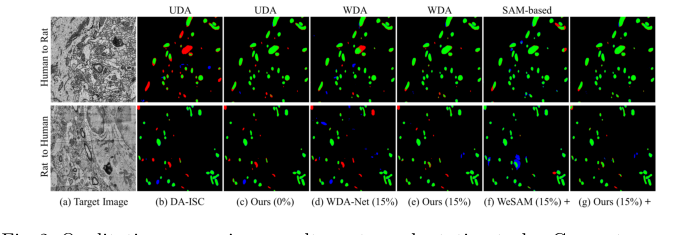 Figure 2. Qualitative comparison results on two adaptation tasks. Green: true pos- itives; Red: false negatives; Blue: false positives
Figure 2. Qualitative comparison results on two adaptation tasks. Green: true pos- itives; Red: false negatives; Blue: false positives
Discussion and Future Evolution
This paper is a brilliant example of how to make foundation models actually useful for specialized scientific domains. However, there are a few areas where we could push this further:
- Source-Free Adaptation: The authors admit their model requires access to the source data. In real-world clinical settings, source data is often proprietary or restricted by privacy laws. Future research could explore "source-free" adaptation, where the model adapts to a new domain using only the target data and the pre-trained weights, without ever seeing the original source images.
- Handling 3D Volumetric Continuity: EM data is often 3D, but this model primarily treats it as a series of 2D images. Integrating temporal or spatial consistency across slices could further reduce the need for sparse points, as the model could "track" an organelle through the volume.
- Uncertainty Quantification: In medical diagnostics, knowing when a model is guessing is as important as the guess itself. Integrating Bayesian uncertainty or conformal prediction into the PCL mechanism could help clinicians identify which segmentations require manual review, making the tool more trustworthy in a lab setting.
