All-in-one medical image-to-image translation
Unifies translation, offers semantic control, and works without fine-tuning.
Background & Academic Lineage
The Origin & Academic Lineage
The problem of medical image-to-image (I2I) translation emerged from the fundamental need to harmonize heterogeneous medical imaging data. In clinical practice, images are acquired using diverse scanning protocols, hardware, and parameter settings, leading to significant "domain shifts" where the statistical distribution of data from one center differs from another. Historically, deep learning models trained on one specific dataset often fail to generalize when applied to data from a different source.
Previous approaches to this problem typically relied on task-specific architectures or discrete labels (like one-hot encoding) to distinguish between imaging domains. These methods suffer from a critical "pain point": they lack scalability. As the number of domains grows, the complexity of managing these discrete labels increases, and the models struggle to capture the subtle, continuous semantic relationships between different imaging sequences. Furthermore, many existing models require target-domain style information during inference, making them unsuitable for real-world clinical deployment where such information is often unavailable. The authors address this by moving away from rigid, discrete labels toward a flexible, prompt-driven framework that leverages natural language to describe imaging protocols, thereby enabling a more robust and generalizable "all-in-one" translation model.
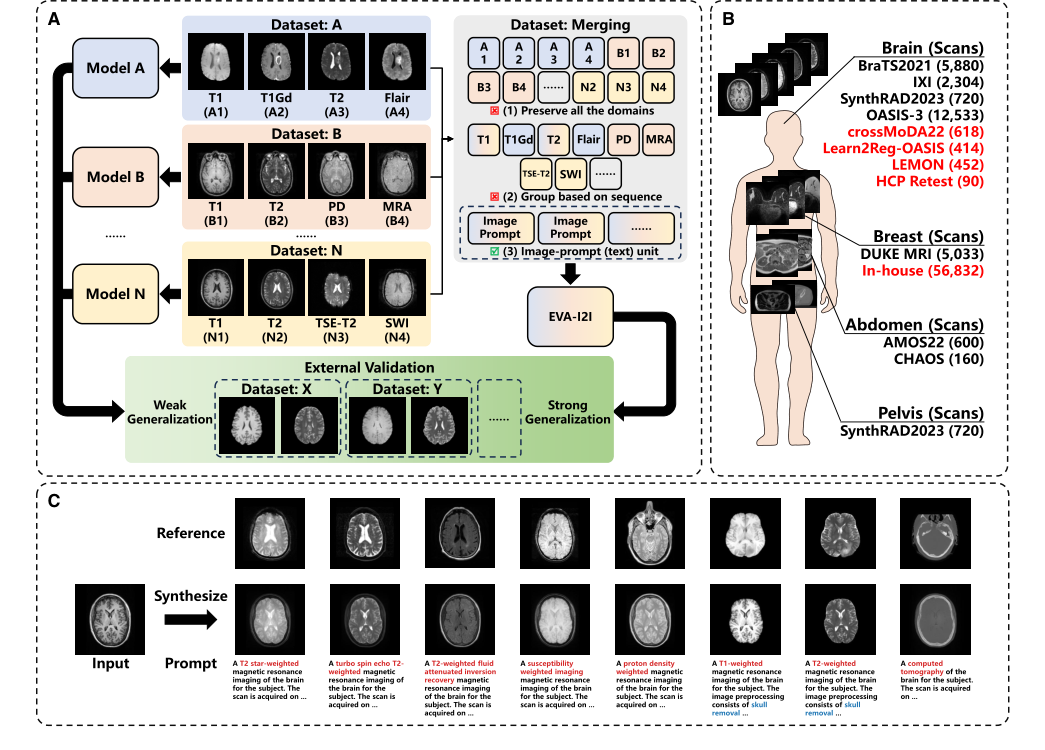 Figure 1. Overview of the proposed EVA-I2I model (A) Integrating domains from different datasets allows training an I2I translation model with strong generalization. (B) Datasets used in this work, where datasets for external validation are in red. (C) The proposed EVA-I2I model transforms the input image to different domains, given prompts. Note that the reference image is an example of the given prompt and is not used for inference
Figure 1. Overview of the proposed EVA-I2I model (A) Integrating domains from different datasets allows training an I2I translation model with strong generalization. (B) Datasets used in this work, where datasets for external validation are in red. (C) The proposed EVA-I2I model transforms the input image to different domains, given prompts. Note that the reference image is an example of the given prompt and is not used for inference
Intuitive Domain Terms
- Domain Shift: Imagine trying to read a book written in a slightly different dialect than the one you learned; you can understand the gist, but the subtle nuances are lost. In medical imaging, this is when a model trained on "Hospital A's" MRI settings struggles to interpret "Hospital B's" images because the contrast or brightness levels are slightly different.
- One-Hot Encoding: Think of this as a rigid filing system where every possible category gets its own unique, isolated box (e.g., "0001" for T1, "0010" for T2). It works for a few items, but if you have hundreds of categories, it becomes impossible to manage, and it fails to show that T1 and T1-contrast images are actually related.
- Contrastive Language-Image Pre-training (CLIP): Imagine a universal translator that has read every medical textbook and looked at every medical scan. It learns to map the text description of a scan (e.g., "T2-weighted brain MRI") to the actual visual patterns in the image, allowing the model to "understand" what a scan should look like based on a text prompt.
- Zero-Shot Adaptation: This is the ability of a model to perform a task on a type of data it has never seen before during its training phase. It is like a doctor who, having studied general anatomy, can successfully diagnose a rare condition they have never personally encountered by applying their broad, foundational knowledge.
Notation Table
| Notation | Description |
|---|---|
| $X$ | The input medical image |
| $P$ | The natural language prompt describing the scan |
| $E_I$ | The image encoder used to extract anatomical features |
| $E_T$ | The text encoder used to process the medical prompt |
| $\Theta_s$ | The domain representation derived from the prompt |
| $G$ | The conditional generator (decoder) for image synthesis |
| $D$ | The DCLIP-driven discriminator |
| $\mathcal{L}_{DCLIP}$ | The loss function for aligning text and image embeddings |
| $\mathcal{L}_{rec}$ | The reconstruction loss for paired training samples |
| $\tau$ | The scalar temperature parameter for contrastive learning |
Problem Definition & Constraints
Core Problem Formulation & The Dilemma
The core problem addressed by this paper is the lack of a scalable and generalizable image-to-image (I2I) translation model for multi-domain medical imaging. Deep learning has shown promise in medical image analysis, but its practical application is severely hampered by the "domain shift" issue. This occurs when training data (source domain) and test data (target domain) come from different distributions, often due to variations in scanning parameters, devices, or even modalities across different clinical centers.
Input/Current State:
The current state involves diverse medical images from multiple domains (e.g., different MRI sequences like T1, T2, Flair, PD, MRA; different organs like brain, breast, abdomen, pelvis; varying scanning protocols and parameters). These images are typically processed by traditional multi-domain I2I approaches that rely on task-specific architectures or discrete domain labels, often using one-hot encoding for domain representation. These methods are limited in their scalability and generalization capabilities. Furthermore, one-to-one I2I translation models face a "geometrical increase" in required models and training time as the number of domains grows, making them impractical for large-scale multi-domain scenarios. Existing models also struggle with external validation, showing a "cynical performance dropping" when applied to new, unseen datasets.
Desired Endpoint (Output/Goal State):
The desired endpoint is a unified, extensible, and generalizable I2I translation framework, named EVA-I2I, that can synthesize medical images across diverse domains simultaneously. This model should effectively encode varied imaging protocols while maintaining semantic consistency and adaptability. The ultimate goal is to enable robust generalization to unseen domains and facilitate direct clinical application without the need for fine-tuning. Specifically, the model aims to synthesize every seen domain at once with a single training session, achieve excellent image quality, and support zero-shot domain adaptation for never-before-seen domains. It should leverage DICOM-tag-informed semantic representations to allow flexible and scalable domain specification, moving beyond the limitations of fixed one-hot codes.
Missing Link or Mathematical Gap:
The exact missing link is a robust and semantically rich mechanism for encoding and controlling diverse medical imaging domains within a unified I2I translation framework. Previous methods, primarily relying on one-hot encoding, fail to capture the subtle semantic relationships between domains (e.g., distinguishing T2 images with skull from those without). This mathematical gap prevents models from scaling effectively to a large number of heterogeneous domains and generalizing to unseen ones while preserving critical anatomical and pathological information. The paper attempts to bridge this by introducing DICOM-tag-informed contrastive language-image pre-training (DCLIP) to create a common latent space that semantically aligns natural language scan descriptions with image features, thereby enabling flexible and scalable domain control.
Painful Trade-off or Dilemma:
The most painful trade-off that has trapped previous researchers is the dilemma between scalability and semantic richness/generalization.
1. Scalability vs. Semantic Richness: Traditional methods using one-hot encoding are simple to implement for a small, fixed number of domains. However, they are inherently unscalable; as the number of medical imaging domains (e.g., different sequences, organs, acquisition parameters) grows, the one-hot encoding vector becomes impractically long, and it fails to capture any semantic relationships between domains. For instance, two T2-weighted images might differ only by the presence or absence of skull, a subtle but semantically important distinction that one-hot encoding cannot represent. This leads to a "geometrical increase" in required models or training time, making multi-domain I2I translation computationally prohibitive and inflexible.
2. Unified Model vs. Task-Specific Performance & Generalization: Developing a single "omnipotent" model capable of translating across all domains simultaneously often comes at the cost of performance on specific, fine-grained tasks or requires extensive fine-tuning for new, external datasets. Previous I2I models, while showing promising results in controlled settings, often exhibit a "cynical performance dropping" when deployed in real-world clinical scenarios with external validation, failing to generalize robustly. This forces researchers to choose between specialized, high-performing but narrow models, or broad, less accurate, and poorly generalizing ones.
Constraints & Failure Modes
This problem is insanely difficult to solve due to several harsh, realistic walls:
Physical Constraints:
* Diverse and Variable Scanning Protocols: Medical images are acquired with a vast array of scanning devices, manufacturers, and subtle parameter settings (e.g., flip angle, echo time, repetition time, fat saturation, contrast agents). These variations create significant domain shifts and make it challenging to integrate datasets and maintain semantic consistency across them.
* Anatomical Complexity: Different organs (brain, breast, abdomen, pelvis) possess vastly different anatomical structures and pathologies. A model designed for brain MRI might struggle with the greater anatomical complexity of abdominal MRI, leading to lower synthesis quality.
* Fixed Image Resolution and 2D Processing: The proposed model is trained with a fixed image resolution (256x256) and is currently only capable of processing 2D slices. This limits its applicability in higher-resolution scenarios and hinders its ability to efficiently comprehend complex 3D structures, potentially leading to inter-slice discontinuities, especially in zero-shot translation tasks.
* Missing Sequences and Small Sample Sizes: In real clinical settings, it is common to encounter small sample sizes or missing sequences for certain target domains. This data sparsity makes direct training of I2I models on these specific domains difficult and often yields poor results.
Computational Constraints:
* Scalability of Model Architectures: Traditional multi-domain I2I models often require the number of generators, encoders, or discriminators to grow linearly or even geometrically with the number of domains. This leads to exponentially increasing computational demands and training times, making them unfeasible for a truly "all-in-one" solution across many diverse domains.
* Hardware Memory Limits: While not explicitly detailed as a hard limit for this paper's solution, the need for efficient model backbones (like ConvNeXt) and the reporting of GFLOPs and inference times imply that computational resources and memory capacity are significant practical constraints in developing and deploying such large-scale models.
Data-Driven Constraints:
* Domain Shift and Lack of Labeled Data: The fundamental challenge is the inherent domain shift between datasets, often caused by varying acquisition parameters across different centers. Compounding this is the lack of a large number of annotated training data that can cover the full spectrum of test set distributions, necessitating robust domain adaptation.
* Limitations of One-Hot Encoding: The prevalent method of using one-hot encoding to represent domains is a major constraint. It cannot distinguish semantic differences between domains (e.g., T2 with skull vs. T2 without skull) and becomes unwieldy and non-scalable as the number of domains expands. This prevents models from learning rich, transferable representations.
* External Validation Performance Drop: I2I models frequently exhibit a significant drop in performance when applied to external validation datasets that were not part of their training distribution. This makes it difficult to develop models that are truly robust and clinically deployable without extensive fine-tuning for each new target domain.
* Inconsistent Out-Domain Coverage: The out-domain datasets used for evaluation may not cover the same organs or imaging domains as the in-domain datasets. This can lead to inconsistencies and hinder direct comparability of performance metrics, making it difficult to accurately assess true generalization capabilities.
* DICOM Tag Availability: While DICOM tags are a rich source of information for prompt generation, for many public datasets, DICOM files are not provided. This necessitates manual collection of relevant information from summary files, which is a laborious and potentially error-prone process.
Why This Approach
The Inevitability of the Choice
The authors arrived at the EVA-I2I framework because traditional methods for multi-domain medical image-to-image (I2I) translation hit a fundamental "scalability wall." Standard approaches, such as one-hot encoding to label domains, fail as the number of domains grows, because the encoding space becomes rigid and lacks the semantic depth required to understand relationships between different imaging protocols. The authors realized that to achieve a truly "omnipotent" model, they needed a representation that could capture the underlying semantics of medical scans rather than treating each domain as an isolated, discrete category. This realization led to the adoption of Contrastive Language-Image Pre-training (CLIP), specifically adapted for DICOM metadata, as the only viable way to map diverse, heterogeneous medical datasets into a unified, continuous latent space.
Comparative Superiority
The superiority of EVA-I2I is rooted in its use of DCLIP (DICOM-tag-informed CLIP) over traditional one-hot or unaligned text encoders like BioBERT. While previous models like MUNIT or StarGAN v.2 rely on style extraction or discrete labels, EVA-I2I leverages the dense, structured semantic information inherent in natural language descriptions of medical scans. Structurally, this allows the model to perform "semantic-driven synthesis," where the generator is conditioned on a rich, aligned text embedding rather than a sparse, non-semantic label. This provides a qualitative advantage: the model can distinguish between subtle imaging differences (e.g., fat saturation vs. non-fat saturation) that one-hot encoding would simply miss or conflate, leading to significantly higher structural fidelity and better generalization to unseen domains.
Alignment with Constraints
The framework is a precise "marriage" between the harsh requirements of clinical medical imaging and the flexibility of vision-language models. The primary constraints—the need for scalability across diverse, multi-center datasets and the requirement for zero-shot generalization—are directly addressed by the DCLIP architecture. By pairing scans with natural language prompts derived from DICOM tags, the authors created a system that does not require explicit domain labels during inference. This allows the model to handle "never-before-seen" domains by simply providing a descriptive prompt, effectively bypassing the need for expensive, task-specific fine-tuning that typically plagues medical AI deployment.
Rejection of Alternatives
The authors explicitly reject one-hot encoding and unaligned text encoders (like BioBERT) because they lack the capacity to capture the complex, hierarchical relationships between medical imaging domains. One-hot encoding was rejected because it is inherently non-extensible and fails to represent the semantic distance between similar but distinct sequences. BioBERT was rejected because, while it understands text, it does not inherently align that text with the visual features of medical images. By contrast, the DCLIP approach ensures that the text and image features are mapped into a shared, cohesive space, which is essential for the model to "understand" what a specific sequence should look like, even if it has never encountered that exact domain during training.
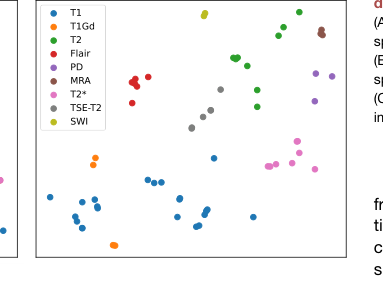 Figure 4. Visualization of prompt embed ding (A) The t-SNE visualization of prompt embedding space of BioBERT encoding. (B) The t-SNE visualization of prompt embedding space of DCLIP encoding. (C) Bar chart of zero-shot classification on medical imaging characters
Figure 4. Visualization of prompt embed ding (A) The t-SNE visualization of prompt embedding space of BioBERT encoding. (B) The t-SNE visualization of prompt embedding space of DCLIP encoding. (C) Bar chart of zero-shot classification on medical imaging characters
Mathematical & Logical Mechanism
The Master Equation
The core mathematical engine of the EVA-I2I framework is the DCLIP loss function, which aligns medical image representations with their corresponding textual prompts. This is defined as:
$$ \mathcal{L}_{\text{DCLIP}} = - \sum_{i \in D} \left( \frac{1}{|\mathcal{M}(i, \cdot)|} \sum_{m \in \mathcal{M}(i, \cdot)} \log \frac{\exp(z_T^{(i)} \cdot z_I^{(m)} / \tau)}{\sum_{j \in D} \exp(z_T^{(i)} \cdot z_I^{(j)} / \tau)} + \frac{1}{|\mathcal{M}(\cdot, i)|} \sum_{m \in \mathcal{M}(\cdot, i)} \log \frac{\exp(z_I^{(i)} \cdot z_T^{(m)} / \tau)}{\sum_{j \in D} \exp(z_I^{(i)} \cdot z_T^{(j)} / \tau)} \right) $$
Term-by-Term Autopsy
- $z_I$ and $z_T$: These are the projected latent representations of the medical image (via the image encoder $E_I$) and the text prompt (via the text encoder $E_T$), respectively. They act as the "semantic coordinates" in a shared embedding space.
- $\mathcal{M}$: This is the confusion matrix (or label matrix) where $\mathcal{M}_{ij} = 1$ if prompt $i$ matches image $j$. It acts as a logical filter to ensure the model learns associations only between valid pairs.
- $\tau$: This is the scalar temperature parameter (set to 0.07). It controls the "sharpness" of the probability distribution; a smaller $\tau$ makes the model more sensitive to small differences in cosine similarity, effectively acting as a contrast booster.
- $\exp(\cdot \cdot \cdot / \tau)$: The exponential function transforms the dot product (cosine similarity) into a positive value, while the division by $\tau$ scales the logits. The author uses this to convert similarity scores into a probability distribution suitable for Softmax-style normalization.
- The summation structure: The author uses a double summation to handle the "many-to-many" nature of medical data, where one prompt (e.g., "T1-weighted brain MRI") might correspond to multiple images from different patients. The division by the cardinality $|\mathcal{M}|$ ensures the loss is normalized, preventing prompts with many associated images from dominating the gradient updates.
Step-by-Step Flow
- Input Encoding: An input image $X$ is passed through the ConvNeXt-based image encoder $E_I$ to produce a vector $z_I$. Simultaneously, a medical prompt $P$ (derived from DICOM tags) is processed by the 12-layer Transformer $E_T$ to yield $z_T$.
- Projection: Both vectors are projected into a common latent space where their semantic alignment is measured.
- Similarity Calculation: The model calculates the dot product between $z_I$ and $z_T$. If the prompt and image are a match, the dot product is maximized.
- Contrastive Normalization: The Softmax-like denominator sums over all other samples in the batch, forcing the model to push non-matching pairs apart in the embedding space while pulling matching pairs together.
- Synthesis: Once aligned, the DCLIP encoding $T$ is fed into the conditional decoder $G$ (using dynamic convolutions) to guide the transformation of the input structure $\Phi$ into the target domain.
Optimization Dynamics
The model learns through a two-stage process. In the initial stage, it uses inner-subject supervision to minimize reconstruction error. In the fine-tuning stage, it incorporates cross-dataset collaborative learning. The loss landscape is shaped by the adversarial loss $\mathcal{L}_{\text{adv}}$, which forces the generator to produce realistic images that the DCLIP-driven discriminator $D$ cannot distinguish from real data. The gradients flow from the DCLIP loss back through the encoders, effectively "sculpting" the latent space so that similar medical protocols cluster together. This ensures that when the model encounters a new, unseen domain, it can map the prompt to a region of the latent space that is already semantically consistent, enabling zero-shot generalization. The use of $\mathcal{L}_{\text{con}}$ and $\mathcal{L}_{\text{tri}}$ acts as a regularizer, preventing the model from collapsing into a trivial solution and ensuring that the structural integrity of the anatomy is preserved during the translation process.
Results, Limitations & Conclusion
Experimental Design & Baselines
To validate the efficacy of the EVA-I2I framework, the authors conducted a comprehensive set of experiments using seven public datasets comprising 27,950 3D medical scans across brain, breast, abdomen, and pelvis domains. The experimental design was structured to test the model's ability to perform multi-domain image-to-image (I2I) translation in a single training session.
The "victims" (baseline models) selected for comparison included established architectures such as MUNIT, StarGAN v2, Chen et al., and Seq2Seq. These baselines were chosen because they represent the state-of-the-art in multi-domain I2I translation. The authors ensured a fair comparison by training all models on the same in-domain training cohorts. To prove the superiority of their DCLIP-driven approach, they performed ablation studies comparing DCLIP against one-hot encoding and BioBERT, as well as evaluating different backbones (ResNet, Swin-Transformer, and the proposed ConvNeXt). The evaluation metrics included Peak Signal-to-Noise Ratio (PSNR), Structural Similarity Index Measure (SSIM), and Learned Perceptual Image Patch Similarity (LPIPS) to quantify both pixel-level fidelity and perceptual quality.
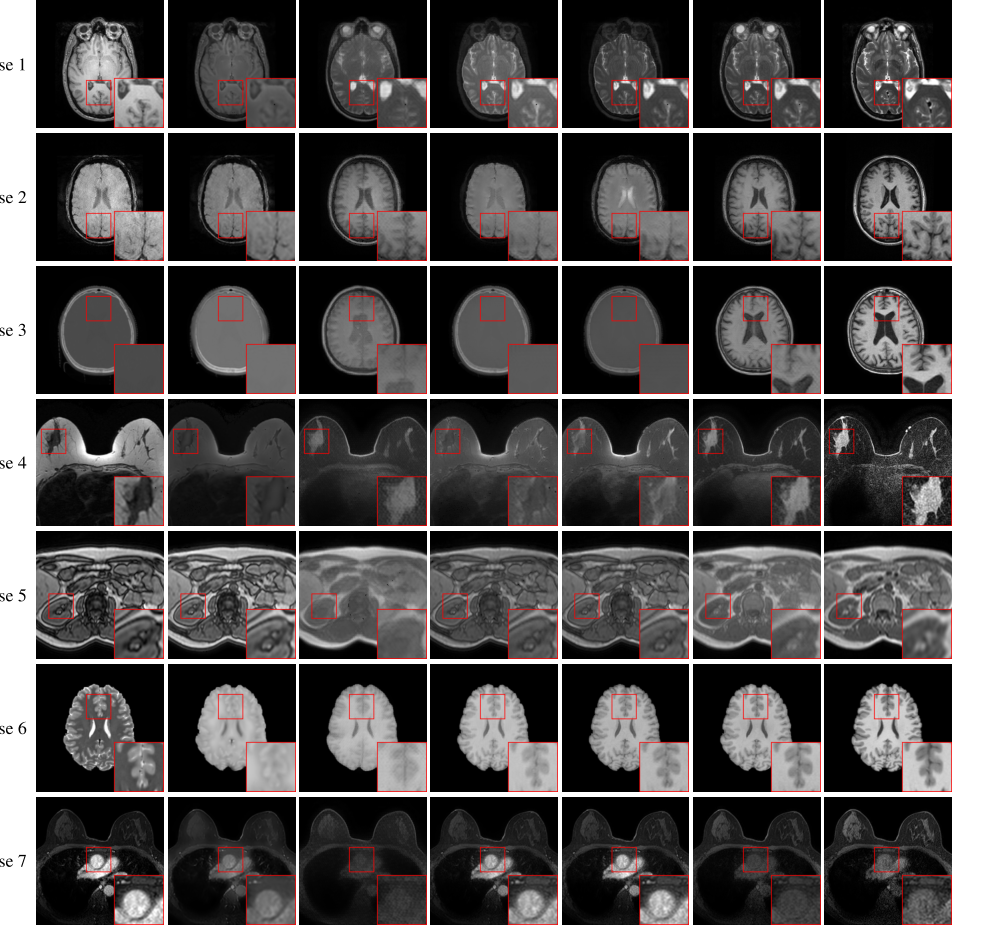 Figure 2. Axial visualization of translated images generated by comparison methods Cases 1–5 from the in-domain datasets show the transformation from T1 to T2, susceptibility weighted imaging (SWI) to T1, CT to T1, non-fat saturated to fat saturated, and out of phase to in phase. Case 6 and case 7 from the out-domain datasets show the transformation from T2 to T1 and post-contrast dynamic contrast-enhanced (DCE) to pre-contrast DCE. Our EVA-I2I attains better quality and a style that resembles the target image
Figure 2. Axial visualization of translated images generated by comparison methods Cases 1–5 from the in-domain datasets show the transformation from T1 to T2, susceptibility weighted imaging (SWI) to T1, CT to T1, non-fat saturated to fat saturated, and out of phase to in phase. Case 6 and case 7 from the out-domain datasets show the transformation from T2 to T1 and post-contrast dynamic contrast-enhanced (DCE) to pre-contrast DCE. Our EVA-I2I attains better quality and a style that resembles the target image
What the Evidence Proves
The evidence provided in the paper is quite compelling. Quantitative results (Table 1) demonstrate that EVA-I2I consistently outperforms the baseline models across both in-domain and out-domain datasets. Specifically, EVA-I2I achieved significantly better synthesis performance ($p < 0.05$ for PSNR and LPIPS) compared to the competition.
The authors provided undeniable evidence of the model's robustness through:
1. Zero-shot Generalization: The model successfully translated images from never-before-seen domains (e.g., wash-in DCE to T1) without requiring fine-tuning, a feat where it surpassed StarGAN v2 in structural detail.
2. Downstream Task Improvement: The model was applied to brain MRI registration, vestibular schwannoma segmentation, and breast cancer pCR prediction. In all cases, EVA-I2I significantly improved the performance of downstream models (e.g., VoxelMorph, nnU-Net) compared to other domain adaptation methods.
3. Semantic Alignment: The t-SNE visualization (Figure 4) clearly shows that DCLIP encoding creates more cohesive feature clusters than BioBERT, proving that the model effectively captures complex domain semantics. The results suggest that the model is not just "memorizing" images but is actually learning a structured, semantic representation of medical imaging protocols.
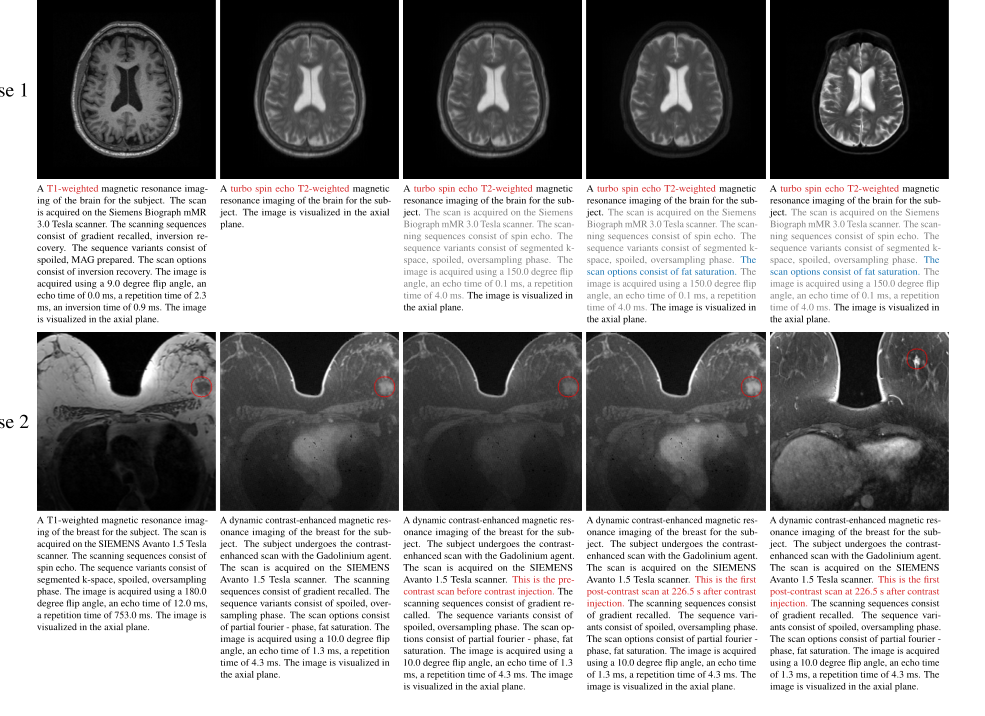 Figure 3. Axial visualization of the impact of changing prompts on generated images In case 1, altering the sequence (prompt 1) results in a noticeable change in the appearance of the image. Including additional scan information in prompt 2 has a minimal impact on the appearance. Fat saturation (prompt 3) further suppresses the intensity of fat tissue under the skin, thus rendering the generated image more consistent with the reference image. In case 2, when the specific time point of DCE-MRI is not specified (prompt 1), the intensity enhancement of the cardiac and tumor (red circle) is between the time points before contrast (prompt 2) and the first post-contrast (prompt 3). Note that the proposed model requires only the input image and the target prompt. The reference image is not used as an input but is provided solely for comparison. It corresponds to the given prompt but is from a different subject because the input case lacks the corresponding image of the reference prompt
Figure 3. Axial visualization of the impact of changing prompts on generated images In case 1, altering the sequence (prompt 1) results in a noticeable change in the appearance of the image. Including additional scan information in prompt 2 has a minimal impact on the appearance. Fat saturation (prompt 3) further suppresses the intensity of fat tissue under the skin, thus rendering the generated image more consistent with the reference image. In case 2, when the specific time point of DCE-MRI is not specified (prompt 1), the intensity enhancement of the cardiac and tumor (red circle) is between the time points before contrast (prompt 2) and the first post-contrast (prompt 3). Note that the proposed model requires only the input image and the target prompt. The reference image is not used as an input but is provided solely for comparison. It corresponds to the given prompt but is from a different subject because the input case lacks the corresponding image of the reference prompt
Limitations & Future Directions
The model is currently restricted to 2D slice processing, which may lead to inter-slice discontinuities when reconstructing 3D volumes. Second, the fixed image resolution used during training limits the model's performance in high-resolution clinical scenarios. Finally, the out-domain datasets used for validation do not fully cover the same organs or imaging domains as the in-domain training set, which can lead to inconsistencies in performance metrics.
Looking forward, I believe there are several exciting avenues for evolution:
* 3D Volumetric Integration: Extending the architecture to process 3D volumes directly would likely resolve the inter-slice inconsistency issues and improve clinical utility.
* Foundation Model Scaling: Given the success of DCLIP, one could explore scaling this to even larger, more diverse datasets, potentially moving toward a truly "universal" medical image translation foundation model.
* Explainability: While the prompts provide a form of control, further research into why the model attends to specific DICOM tags could provide clinicians with more confidence in the synthesized results, making the "black box" of AI more transparent.
These findings suggest that we are moving toward a future where a single, generalizable model can handle the vast heterogeneity of medical imaging, reducing the need for tedious, task-specific fine-tuning.
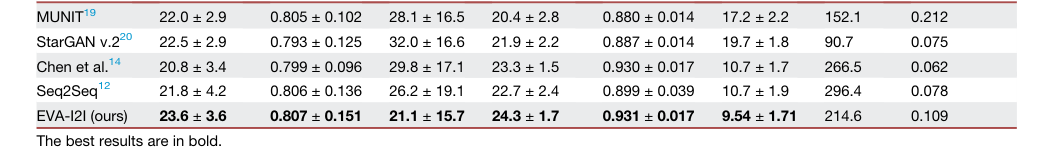 Table 2. illustrates the quantitative results of the ablation study on domain encoding, model backbone, and training loss function. We compare three domain encodings to equip our EVA-I2I, including one hot,12 BioBERT,35 and DCLIP (proposed). BioBERT encoding can extract semantic information from medical prompts, leading to better synthesis results than one- hot encoding, and our proposed DCLIP encoding aligns text and images, making its synthesis performance significantly (p < 0:05 for PNSR and LPIPS) better than BioBERT encoding, which cannot align text and images. We also compare three model backbones of the structure encoder EΦ in EVA-I2I, including ResNet,36 Swin-Transformer,37 and ConvNeXt (pro posed). The ConvNeXt backbone slightly outperforms other methods. Moreover, we also evaluate the effectiveness of cross-dataset collaborative learning. Results show that the model trained with Lfine significantly (p < 0:05 for PNSR and LPIPS) outperforms the model only trained with Linit
Table 2. illustrates the quantitative results of the ablation study on domain encoding, model backbone, and training loss function. We compare three domain encodings to equip our EVA-I2I, including one hot,12 BioBERT,35 and DCLIP (proposed). BioBERT encoding can extract semantic information from medical prompts, leading to better synthesis results than one- hot encoding, and our proposed DCLIP encoding aligns text and images, making its synthesis performance significantly (p < 0:05 for PNSR and LPIPS) better than BioBERT encoding, which cannot align text and images. We also compare three model backbones of the structure encoder EΦ in EVA-I2I, including ResNet,36 Swin-Transformer,37 and ConvNeXt (pro posed). The ConvNeXt backbone slightly outperforms other methods. Moreover, we also evaluate the effectiveness of cross-dataset collaborative learning. Results show that the model trained with Lfine significantly (p < 0:05 for PNSR and LPIPS) outperforms the model only trained with Linit
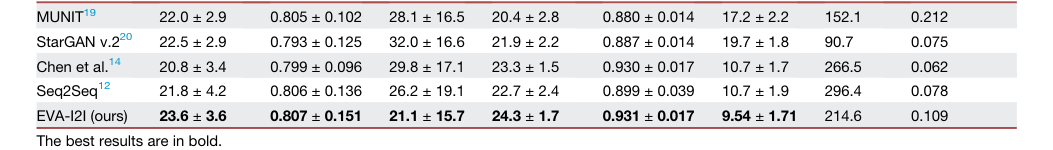 Table 1. The quantitative results of image translation for different comparisons on in- and out-domain datasets
Table 1. The quantitative results of image translation for different comparisons on in- and out-domain datasets
