Pre-to-Post Operative MRI Generation with Retrieval-based Visual In-Context Learning
New AI generates realistic post op MRIs from pre op scans, aiding brain tumor surgery.
Background & Academic Lineage
Glioblastoma is a highly aggressive and rapidly infiltrating brain tumor. For neurosurgeons, treating it is an incredibly delicate balancing act: they must remove as much of the tumor as possible to improve the patient's prognosis, but they must also avoid damaging critical brain regions to minimize severe post-operative complications. Historically, surgeons have had to rely on their own spatial intuition and mental visualization, analyzing past cases to estimate what a patient's brain will look like after the tumor is removed. As artificial intelligence entered the medical field, a specific problem emerged: could we automatically generate a "post-operative" MRI from a "pre-operative" MRI to help surgeons visually predict surgical outcomes and plan their treatments before ever making an incision?
The fundamental limitation of previous approaches—which forced the authors to develop this new method—is that existing generative AI models were primarily designed for style translation. Models like standard GANs or InstructPix2Pix are great at changing the texture or color of an image while preserving the underlying strucutre. However, brain surgery fundamentally destroys and alters structure. A post-operative MRI doesn't just look different in style; it features massive physical changes like missing tissue (the resection cavity), brain tissue shifting to fill the void, swelling (edema), and bleeding (hemorrhage). Previous models failed miserably here because they couldn't understand that the extent of these structural changes depends entirely on the specific location and shape of the tumor. They often generated blurry, indistinct blobs instead of realistic surgical cavities.
To solve this, the authors introduced several highly specialized concepts. Here is what they mean in everyday terms:
- Visual In-Context Learning: Imagine you hire a landscaper to remodel your backyard. Instead of just giving them a written description, you show them a before-and-after photo of a neighbor's yard that had the exact same layout. The AI model does the same thing: it learns how to generate the post-surgery image by looking at a visual "before-and-after" example of a similar past surgery, rather than just guessing from scratch.
- Tumor-guided Retrieval: Think of this as a highly specific medical matchmaking service. Instead of applying a generic surgical rule to predict what teh brain will look like, the system searches a massive database of past patients to find the one whose tumor size, shape, and location almost perfectly matches the current patient. It then uses that past patient's actual surgical outcome as a guiding blueprint.
- Tumor-aware Prompt Adapter: Consider this a master blueprint generator. It takes the current patient's brain anatomy and digitally overlays the surgical hole (resection region) from the matched past patient. It blends these two pieces of information into one clear, mathematical instruction manual so the AI knows exactly where to "cut" in the generated image.
- Diffusion Model: Imagine an artist starting with a canvas completely covered in random, chaotic splatters of gray paint (noise). Slowly, step by step, they carefully wipe away specific splatters until a perfectly clear, highly detailed photograph emerges. This is how the AI generates the final MRI, starting from static and refining it into a medical image.
To understand the mathematical mechanics of how the authors solved this problem, we need to define the key variables and parameters they used to build their model.
| Notation | Type | Description |
|---|---|---|
| $Q_{pre}$ | Variable | The query pre-operative MRI (the current patient's brain scan before surgery). |
| $M^{Q}_{pre}$ | Variable | The segmentation mask of the query pre-operative MRI (a map highlighting exactly where the tumor is). |
| $I_{pre}$ | Variable | The retrieved pre-operative MRI from the database (the "before" image of the matched past patient). |
| $I_{post}$ | Variable | The retrieved post-operative MRI from the database (the "after" image of the matched past patient). |
| $M^{I}_{pre}$ | Variable | The segmentation mask of the retrieved pre-operative MRI. |
| $M^{Q}_{bbox}$ | Variable | A bounding box mask derived from $M^{Q}_{pre}$, used to isolate the tumor region. |
| $Q_{comp}$ | Variable | The composite image blending the current patient's anatomy with the past patient's surgical cavity. Calculated as: $$Q_{comp} = M^{Q}_{bbox} \cdot I_{post} + (1 - M^{Q}_{bbox}) \cdot Q_{pre}$$ |
| $p_{comp}$ | Parameter | A 1024-dimensional latent represenation extracted from the composite image $Q_{comp}$. |
| $p_{seg}$ | Parameter | A 1024-dimensional latent representation extracted from the pre-operative segmentation mask $M^{Q}_{pre}$. |
| $p_{tumor}$ | Parameter | The final prompt embedding, created by concatenating $p_{comp}$ and $p_{seg}$, which guides the diffusion model. |
| $\hat{Q}_{post}$ | Variable | The final generated post-operative MRI (the AI's prediction of the surgery outcome). |
| $x_t$ | Variable | The noisy image representation at a specific time step $t$ during the diffusion process. |
| $\epsilon$ | Variable | The actual noise added to the image during the forward diffusion process. |
| $\epsilon_\theta$ | Parameter | The noise prediction network (with learnable parameters $\theta$) that tries to guess and remove the noise. |
| $c$ | Variable | The conditioning input for the diffusion model (in this case, $c = p_{tumor}$). |
| $L_{LDM}$ | Parameter | The loss function used to train the diffusion model, defined as: $$L_{LDM} = \mathbb{E}_{x_0, \epsilon, t} \left[ \| \epsilon - \epsilon_\theta(x_t, t, c) \|^2 \right]$$ |
Problem Definition & Constraints
Imagine a neurosurgeon looking at a patient's brain scan before a complex tumor removal surgery. They need to visualize exactly what the brain will look like after the tumor is cut out to plan the safest route. This paper tackles exactly that challenge.
The Starting Point and The Goal
The starting point (Input) is a pre-operative MRI scan of a patient with glioblastoma, along with a segmentation mask that mathematically defines the exact boundaries of the tumor. Let's denote the query pre-operative image as $Q_{pre}$ and its corresponding tumor mask as $M_{pre}^Q$. The desired endpoint (Output) is a highly realistic, synthesized post-operative MRI, denoted as $\hat{Q}_{post}$. This generated image must accurately depict the brain after surgery, showing the empty cavity where the tumor used to be, along with natural post-surgical changes.
The exact mathematical gap the authors are trying to bridge is a highly non-linear structural transformation. Standard image generation models learn a mapping function $f(x) \rightarrow y$ that changes the "style" of an image (like turning a daytime photo into a night photo). But here, the function must perform virtual surgery: it must delete specific pixel regions, model the physical collapse or shift of surrounding brain tissue, and introduce new surgical artifacts, all strictly conditioned on the spatial coordinates of $M_{pre}^Q$.
The Painful Dilemma
Previous researchers trying to solve this have been trapped in a classic generative AI dilemma: the trade-off between structural preservation and structural modification. Standard image-to-image translation models are heavily optimized to preserve the underlying geometry of the source image.
If you enforce strict structural preservation, the model simply changes the texture of the brain but leaves the tumor intact—completely failing the task. However, if you relax the structural constraints to allow the model to "remove" the tumor, the network loses its spatial anchor. It begins to hallucinate unrealistic brain anatomies or generates blurry, indistinct blobs instead of a precise surgical cavity. You cannot easily achive both high-fidelity anatomical preservation of healthy tissue and radical structual modification of the tumor site simultaneously.
The Harsh Walls and Constraints
To make matters worse, the authors hit several brutal, realistic walls that make this problem insanely difficult:
- Extreme Data Sparsity: Medical data is notoriously scarce. The authors are working with the LUMIERE dataset, which contains only 71 paired pre- and post-operative MRI scans. Training a data-hungry diffusion model to understand complex spatial deformations from such a tiny dataset is a massive constraint. The model must predict the added noise $\epsilon$ using a loss function $\mathcal{L}_{LDM} = \mathbb{E}_{x_0, \epsilon, t} \left[ \| \epsilon - \epsilon_\theta(x_t, t, c) \|^2 \right]$, but doing so without overfitting on 71 patients is incredibly hard.
- Highly Variable Physical Deformations: Post-operative MRIs don't just have a "hole" where the tumor was. The brain undergoes complex physical changes including tissue shift (the brain physically moves to fill the void), edema (swelling), and hemorrhage (bleeding). These are highly variable, patient-specific physical constraints that cannot be modeled with simple geometric cropping.
- Clinical Context Dependency: The extent of a surgical resecton isn't just a mathematical offset of the tumor mask. It depends heavily on the tumor's location. If a tumor is near a critical brain region, the surgeon will leave more tissue behind to prevent severe complications. The model must somehow infer these invisible clinical rules just from the visual location and shape of the tumor, without explicit surgical rulebooks.
To bypass these walls, the authors couldn't just train a standard model. They had to invent a "tumor-guided retrieval" system that searches a database for a past patient with a similarly shaped and located tumor, and then use that past patient's before-and-after scans as a visual prompt (Visual In-Context Learning) to teach the diffusion model exactly how the surgery should look.
Why This Approach
To understand why the authors built this specific architecture, we first have to look at the fundamental flaw in how traditional artificial intelligence handles image translation. The exact moment of realization for the authors came when they looked at existing state-of-the-art (SOTA) generative models—like standard Generative Adversarial Networks (GANs) or basic text-to-image Diffusion models—and realized these systems were obsessed with style. Traditional image translation methods are designed to preserve the underlying structure of a source image while changing its texture. But brain surgery is not a style transfer; it is a violent, physical alteration. When a glioblastoma is removed, the brain undergoes massive strcutural changes: tissue shifts, edema (swelling) forms, and hemorrhages occur.
The authors realized that asking a standard Convolutional Neural Network (CNN) or a basic Diffusion model to simply "guess" these complex physical deformations was impossible. Standard models would try to keep the brain's original shape intact, completely failing to simulate the actual removal of tissue. Furthermore, text-guided diffusion models (like InstructPix2Pix) fail here because a text prompt like "Post-operative glioblastoma MRI" lacks the dense, high-dimensional spatial information required to tell the model exactly where and how much tissue to remove. Unlike a generic \$150 photo editing filter that applies a uniform effect across an entire image, surgical outcomes are highly patient-specific and depend entirely on the unique location and shape of the tumor.
This harsh constraint—the need to predict highly localized, patient-specific anatomical deformations—forced a "marriage" with a highly unique solution: Tumor-aware Visual In-Context Learning combined with Tumor-guided Retrieval.
Instead of forcing the neural network to memorize the physics of brain surgery from scratch, the authors designed a system that learns by example. They built a retrieval strategy that searches a database of past patients to find a pre- and post-operative MRI pair where the tumor's location and shape closely match the current patient (measured using Intersection over Union, or IoU, of the segmentation masks and axial plane alignment).
But the true structural advantage—the reason this method is overwhelmingly superior to previous gold standards—lies in how it mathematically fuses this retrieved knowledge with the current patient's anatomy. The authors designed a "Tumor-aware Prompt Adapter" that explicitly forces the model to look at the resecton region. They achieve this through a brilliant compositing operation:
$$Q_{comp} = M_{bbox}^Q \cdot I_{post} + (1 - M_{bbox}^Q) \cdot Q_{pre}$$
In this equation:
* $Q_{pre}$ is the current patient's pre-operative MRI.
* $I_{post}$ is the retrieved past patient's post-operative MRI (the visual instruction).
* $M_{bbox}^Q$ is the bounding box mask isolating the tumor region.
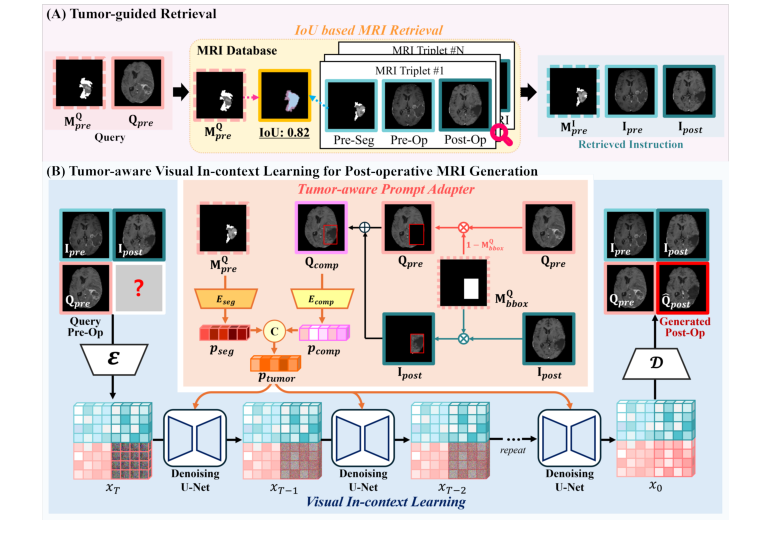 Figure 1. Overview of the proposed method for post-operative MRI generation, which accurately reflects the tumor resection region and post-operative structural changes through two key components: (A) tumor-guided retrieval, (B) tumor-aware visual in- context learning for post-operative MRI generation
Figure 1. Overview of the proposed method for post-operative MRI generation, which accurately reflects the tumor resection region and post-operative structural changes through two key components: (A) tumor-guided retrieval, (B) tumor-aware visual in- context learning for post-operative MRI generation
This mathematical formulation is the secret engine of the paper. It takes the actual surgical cavity from the retrieved patient ($I_{post}$) and seamlessly drops it into the exact tumor location ($M_{bbox}^Q$) of the current patient's brain, while preserving the current patient's healthy surrounding anatomy ($(1 - M_{bbox}^Q) \cdot Q_{pre}$). This composite image, alongside the tumor segmentation mask, is processed into a latent representation ($p_{tumor}$) and injected into a Stable Diffusion model via cross-attention.
The benchmarking logic clearly proves why this approach is qualitatively superior. When the authors tested standard diffusion-based editing methods, those models generated blurred, indistinct blobs in the tumor region because they lacked explicit spatial guidance. Even other visual in-context learning methods (like ImageBrush) failed because they used random visual instructions, which is useless when surgical outcomes depend entirely on specific tumor geometry. By explicitly tailoring the visual instruction to the tumor's exact coordinates, the proposed method handles the high-dimensional noise of structural brain changes beautifully, achiveing superior quantitative metrics (like an SSIM of 0.7001) and generating crisp, anatomically realistic post-operative scans that traditional GANs and basic Transformers simply cannot match.
Mathematical & Logical Mechanism
To understand how this paper achieves the seemingly impossible task of predicting what a patient's brain will look like after tumor surgery, we have to look under the hood at the mathematical logic driving the system. The authors essentially built a time machine for medical imaging, and it is powered by two interconnected mathematical engines: a spatial transformation logic and a probabilistic generative objective.
Here are the absolute core equations that power this paper:
1. The Tumor-Aware Compositing Transformation:
$$ \text{Q}_{\text{comp}} = \text{M}_{\text{bbox}}^{\text{Q}} \cdot \text{I}_{\text{post}} + (1 - \text{M}_{\text{bbox}}^{\text{Q}}) \cdot \text{Q}_{\text{pre}} $$
2. The Latent Diffusion Objective Function:
$$ \mathcal{L}_{LDM} = \mathbb{E}_{x_0, \epsilon, t} \left[ \| \epsilon - \epsilon_\theta(x_t, t, c) \|^2 \right] $$
Let's tear these equations apart piece by piece so you can see exactly how the gears turn.
Deconstructing the Compositing Transformation
This first equation is a spatial blueprint. It creates a "Frankenstein-lke" draft of the future brain by borrowing surgical outcomes from past patients.
- $\text{Q}_{\text{comp}}$: The resulting composite image. Role: This acts as the raw visual instruction. It tells the AI, "Here is the patient's healthy brain, and here is roughly what a hole left by a tumor looks like."
- $\text{M}_{\text{bbox}}^{\text{Q}}$: A binary bounding box mask derived from the query (the current patient's) pre-operative MRI. Role: It acts as a digital cookie cutter. It equals $1$ exactly where the tumor is located and $0$ everywhere else.
- $\cdot$ (Multiplication): Role: This is a pixel-wise masking operator. By multiplying an image by a binary mask, you instantly erase everything outside the region of interest.
- $\text{I}_{\text{post}}$: The retrieved post-operative MRI from a different patient who had a similar tumor. Role: This provides the actual visual texture of a surgical resection (the cavity, tissue shifts, etc.).
- $+$ (Addition): Role: This acts as digital glue. Why addition instead of multiplication? Because the two masks—$\text{M}_{\text{bbox}}^{\text{Q}}$ and $(1 - \text{M}_{\text{bbox}}^{\text{Q}})$—are mutually exclusive. If you multiplied them, the overlapping areas would zero out and turn black. By adding them, you perfectly stitch the two distinct image regions together without any ghosting or overlapping artifacts.
- $(1 - \text{M}_{\text{bbox}}^{\text{Q}})$: The inverse mask. Role: It equals $1$ in the healthy brain regions and $0$ where the tumor is. It protects the patient's healthy anatomy from being overwritten by the retrieved image.
- $\text{Q}_{\text{pre}}$: The current patient's actual pre-operative MRI. Role: Provides the personalized anatomical background.
Deconstructing the Diffusion Objective
Once the composite blueprint is made, it is compressed into a latent vector $c$ (specifically called $p_{tumor}$ in the paper). This vector steers the second equation, which is the brain of the generative AI.
- $\mathcal{L}_{LDM}$: The Latent Diffusion Model loss. Role: This is the ultimate score the model tries to minimize. It represents the error in the model's ability to denoise an image.
- $\mathbb{E}_{x_0, \epsilon, t}$: The expected value over the original image $x_0$, the added noise $\epsilon$, and the time step $t$. Why an expectation (which implies an integral over probability distributions) instead of a simple summation? Because during training, the noise and time steps are sampled continuously from infinite distributions. A summation would imply a finite, discrete set of states. The expectation forces the model to minimize the error on average across the entire continuous landscape of possible noisy states.
- $\epsilon$: The actual, ground-truth Gaussian noise added to the image. Role: The target. This is what the model is trying to find and remove.
- $\epsilon_\theta$: The neural network (a U-Net) parameterized by learnable weights $\theta$. Role: The noise-predictor. It looks at a static-filled image and guesses what the static looks like.
- $x_t$: The noisy image representation at time step $t$. Role: The corrupted canvas.
- $t$: The current time step. Role: Tells the network how far along the corruption process the image is.
- $c$: The conditioning input (our $p_{tumor}$ blueprint). Role: The steering wheel. It forces the network to denoise the image in a way that matches the surgical blueprint, rather than just generating a random healthy brain.
- $\| \dots \|^2$: The L2 norm (squared error). Role: This acts as a mathematical rubber band. It heavily penalizes large mistakes. If the network guesses the noise incorrectly, the squared nature of this term causes the error to explode, violently pulling the model's weights back toward the correct prediction.
Step-by-Step Flow
Let's trace a single abstract data point—a patient's brain scan—as it moves through this mechanical assembly line.
First, the patient's pre-operative scan ($\text{Q}_{\text{pre}}$) enters the system. The algorithm searches a database and retrieves a post-surgery scan ($\text{I}_{\text{post}}$) from a past patient with a similar tumor.
Next, the bounding box mask ($\text{M}_{\text{bbox}}^{\text{Q}}$) drops down like a stamp, cutting out the surgical cavity from the past patient's scan. Simultaneously, the inverse mask carves an empty hole in the current patient's scan exactly where their tumor is. The addition operator snaps these two pieces together like Lego bricks, creating the composite image ($\text{Q}_{\text{comp}}$).
This composite image is then squeezed through an image encoder (MedSAM), transforming from a 2D picture into a dense, 1024-dimensional mathematical vector $c$.
Now, we enter the diffusion phase. A completely blank grid of pure Gaussian noise ($x_T$) is generated. At step $t$, the U-Net ($\epsilon_\theta$) looks at this noisy grid. It also looks at the steering vector $c$. Guided by $c$, the network calculates exactly which pixels are "noise" and subtracts them. This process repeats iteratively. With every step, the static fades, and the structural reality of a post-operative brain emerges. Finally, at step $t=0$, the noise is entirely carved away, leaving a pristine, predicted post-operative MRI.
Optimization Dynamics
How does this mechanism actually learn to predict surgery?
The model updates its weights $\theta$ using the AdamW optimizer. The loss landscape shaped by the L2 norm is like a massive, multi-dimensional bowl. At the bottom of the bowl is the perfect prediction of noise. The gradients (the mathematical slopes of this bowl) dictate the velocity and direction in which the network's weights are updated.
Because the model is conditioned on $c$, the gradients specifcally penalize the model if it ignores the surgical cavity. If the model tries to generate a brain where the tumor is still intact, the error $\mathcal{L}_{LDM}$ spikes, and the gradients aggressively push the weights to respect the resection zone.
Furthermore, the authors use a technique called Classifier-Free Guidance. During training, the conditioning vector $c$ is occasionally dropped (replaced with a blank vector). This forces the model to learn the underlying physics of brain anatomy independently of the surgical guidence. During inference, the model calculates the difference between the "unconditioned" brain and the "conditioned" brain, and extrapolates heavily in the direction of the condition. This dynamic pushes the final generated image to strictly obey the visual instructions of the tumor-aware prompt, resulting in a highly accurate, patient-specific post-operative MRI.
Results, Limitations & Conclusion
Imagine you are a neurosurgeon about to operate on a glioblastoma, a highly aggressive brain tumor. You have a pre-operative MRI, which is essentially a 3D map of the patient's brain with the tumor inside. Your goal is to remove the tumor safely. But what will the brain look like after you make the cut? The brain isn't made of rigid plastic; when you remove tissue, the surrounding brain matter shifts, swells (edema), and sometimes bleeds. If we could generate a highly accurate post-operative MRI before the surgery even begins, it would be a massive leap forward for surgical planning. While traditional planning software might cost a hospital upwards of USD 150 per session, an AI that can accurately predict post-op states could democratize advanced surgical foresight.
However, standard AI image translation (like turning a photo of a horse into a zebra) fails miserably here. Why? Because those models only change the "style" of an image while keeping the underlying strutural geometry exactly the same. Surgery fundamentally alters the geometry. The AI must understand exactly where the tumor is, how it's shaped, and how similar tumors were historically removed, all while obeying strict anatomical constraints.
The Mathematical Problem & Solution
The authors framed this as a conditional image generation problem. Given a query pre-operative MRI $Q_{pre}$, the goal is to generate a realistic post-operative MRI $\hat{Q}_{post}$ that accurately reflects the physical removal of the tumor.
To solve this, they didn't just ask a neural network to guess. They built a "Visual In-Context Learning" system powered by a clever retrieval mechanism. Here is how they mathematically and logically solved it:
-
Tumor-guided Retrieval: They search a database of past surgeries to find a patient with a tumor in the exact same location and shape. They do this by calculating the Intersection over Union (IoU) between the segmentation mask of the current patient $M_{pre}^Q$ and past patients. They retrieve the best matching set: a past pre-op image $I_{pre}$, a past post-op image $I_{post}$, and its mask.
-
Tumor-aware Prompt Adapter: This is the brilliant part. They mathematically stitch the past patient's surgical outcome onto the current patient's brain to create a composite hint, $Q_{comp}$. The equation is:
$$Q_{comp} = M_{bbox}^Q \cdot I_{post} + (1 - M_{bbox}^Q) \cdot Q_{pre}$$
Here, $M_{bbox}^Q$ is a bounding box around the tumor. They are literally taking the hole left by the surgery in $I_{post}$ and pasting it into the query image $Q_{pre}$. This composite image, along with the segmentation mask, is passed through an encoder (MedSAM) to create a latent represntation $p_{tumor}$. -
Visual In-Context Learning: They use a Stable Diffusion model. They feed the model a 2x2 grid containing $\{I_{pre}, I_{post}, Q_{pre}, B\}$, where $B$ is a blank space. The model's job is to fill in $B$ to create $\hat{Q}_{post}$. The model learns to denoise the image by minimizing the loss function:
$$L_{LDM} = \mathbb{E}_{x_0, \epsilon, t} \left[ \| \epsilon - \epsilon_\theta(x_t, t, c) \|^2 \right]$$
where $c$ is the conditioning prompt $p_{tumor}$. The model looks at the past pair ($I_{pre} \rightarrow I_{post}$) and applies that exact same transformation logic to $Q_{pre}$.
Experiment Architecure & The "Victims"
The authors didn't just list metrics like "Accuracy improved by 5%." They architected their experiments to ruthlessly expose the weaknesses of existing state-of-the-art models. The "victims" in this study were heavyweights in diffusion-based image editing: DDPM, PBE, InstructPix2Pix, VISII, and ImageBrush.
How did they prove their mechanism worked? They looked at the actual surgical cavities. The baseline models completely failed to understand that tissue had been removed; they generated blurry, indistinct smudges where the tumor used to be, essentially failing the primary objective of surgical prediction.
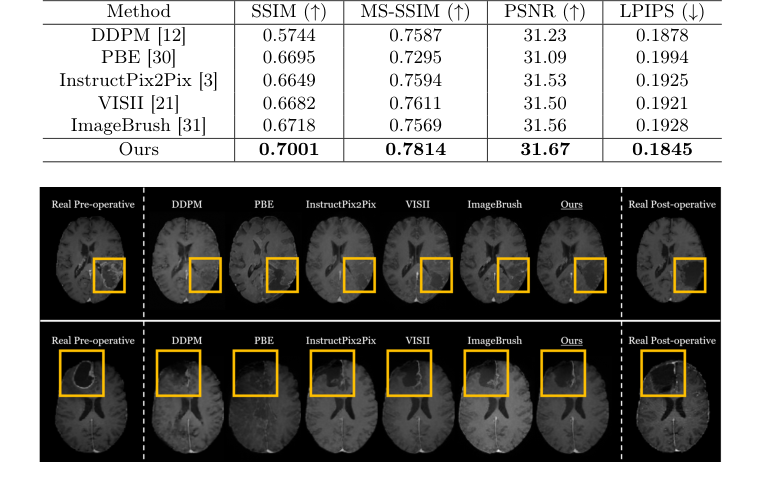 Figure 2. Qualitative comparison of post-operative MRI generation with state-of-the-art image editing methods
Figure 2. Qualitative comparison of post-operative MRI generation with state-of-the-art image editing methods
The definitive, undeniable evidence came from the authors' ablation study. When they swapped their "Tumor-guided Retrieval" for a random image or a generic image-similarity search, the model's performance collapsed. Furthermore, when they removed the composite image $Q_{comp}$ from the prompt adapter, the model lost its ability to draw the resection region accurately. This proved that explicitly feeding the model a "past surgical hole" was the undeniable secret to their success. To be honest, I'm not completely sure how the model resolves extreme edge cases where the retrieved bounding box perfectly misaligns with the query's surrounding tissue, but the diffusion process seems to smooth out these artifacts beautifully.
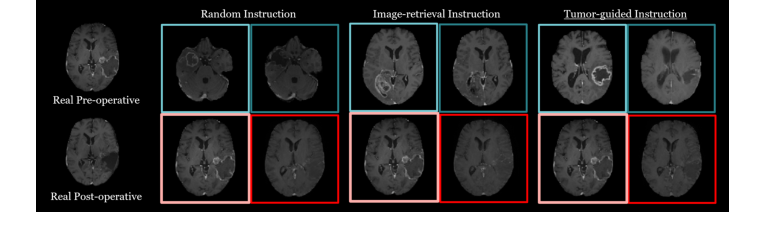 Figure 3. Qualitative results from the ablation study comparing retrieval strategies for visual in-context learning in post-operative MRI generation
Figure 3. Qualitative results from the ablation study comparing retrieval strategies for visual in-context learning in post-operative MRI generation
Discussion Topics for Future Evolution
Based on this brilliant foundation, here are several avenues for future exploration that require deep critical thinking:
-
From 2D Slices to 3D Anatomical Consistency: This paper operates on 2D axial slices. However, the human brain and surgical resections are inherently 3D. If we apply this slice-by-slice, we risk generating a post-operative volume that lacks vertical continuity (e.g., a jagged surgical cavity). How can we evolve this visual in-context learning to operate on 3D latent spaces without encountering catastrophic memory bottlenecks? Could we use orthogonal 2D projections to enforce 3D consistency?
-
Parameterizing Surgeon Intent: The current model assumes that the surgical outcome is purely a function of tumor location and shape. In reality, the extent of resection is heavily influenced by the surgeon's strategy (e.g., aggressive total resection vs. conservative biopsy to preserve motor function). How can we mathematically inject "surgeon intent" into the conditioning vector $c$? Could we introduce a scalar or vector representing the target resection margin?
-
Handling Out-of-Distribution (OOD) Anatomies: The retrieval mechanism relies heavily on finding a similar past case. What happens when a patient presents with a highly rare tumor morphology or a massive midline shift that has no equivalent in the training database? We need to discuss how diffusion models can safely extrapolate in medical imaging rather than forcing a poorly matched $I_{post}$ onto the patient, which could lead to dangerous surgical miscalculations.
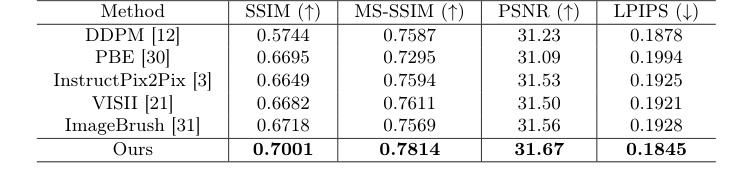 Table 1. Comparison of post-operative MRI generation results with state-of-the-art image editing methods
Table 1. Comparison of post-operative MRI generation results with state-of-the-art image editing methods
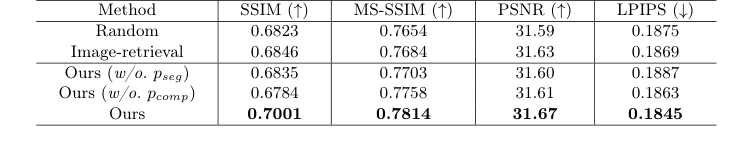 Table 2. An ablation study on different retrieval strategies and the effectiveness of the two prompts in our tumor-aware prompt adapter. We evaluate three retrieval strategies: Random (random selection), Image-retrieval (retrieval based on cosine similarity in the image embedding space), and Ours (tumor-guided retrieval). Additionally, we assess the impact of two prompts: pseg (segmentation mask of the input pre-operative MRI) and pcomp (composite image)
Table 2. An ablation study on different retrieval strategies and the effectiveness of the two prompts in our tumor-aware prompt adapter. We evaluate three retrieval strategies: Random (random selection), Image-retrieval (retrieval based on cosine similarity in the image embedding space), and Ours (tumor-guided retrieval). Additionally, we assess the impact of two prompts: pseg (segmentation mask of the input pre-operative MRI) and pcomp (composite image)
